Causal Trees/Forests Treatment Effects Estimation and Tree Visualization¶
[1]:
import pandas as pd
import numpy as np
import multiprocessing as mp
from collections import defaultdict
np.random.seed(42)
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
import causalml
from causalml.metrics import plot_gain, plot_qini, qini_score
from causalml.dataset import synthetic_data
from causalml.inference.tree import plot_dist_tree_leaves_values, get_tree_leaves_mask
from causalml.inference.meta import BaseSRegressor, BaseXRegressor, BaseTRegressor, BaseDRRegressor
from causalml.inference.tree import CausalRandomForestRegressor
from causalml.inference.tree import CausalTreeRegressor
from causalml.inference.tree.plot import plot_causal_tree
import matplotlib.pyplot as plt
import seaborn as sns
%config InlineBackend.figure_format = 'retina'
Using `tqdm.autonotebook.tqdm` in notebook mode. Use `tqdm.tqdm` instead to force console mode (e.g. in jupyter console)
Failed to import duecredit due to No module named 'duecredit'
[2]:
import importlib
print(importlib.metadata.version('causalml') )
0.14.0
[3]:
# Simulate randomized trial: mode=2
y, X, w, tau, b, e = synthetic_data(mode=2, n=15000, p=20, sigma=5.5)
df = pd.DataFrame(X)
feature_names = [f'feature_{i}' for i in range(X.shape[1])]
df.columns = feature_names
df['outcome'] = y
df['treatment'] = w
df['treatment_effect'] = tau
[4]:
df.head()
[4]:
| feature_0 | feature_1 | feature_2 | feature_3 | feature_4 | feature_5 | feature_6 | feature_7 | feature_8 | feature_9 | ... | feature_13 | feature_14 | feature_15 | feature_16 | feature_17 | feature_18 | feature_19 | outcome | treatment | treatment_effect | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.496714 | -0.138264 | 0.358450 | 1.523030 | -0.234153 | -0.234137 | 1.579213 | 0.767435 | -0.469474 | 0.542560 | ... | -1.913280 | -1.724918 | -0.562288 | -1.012831 | 0.314247 | -0.908024 | -1.412304 | 7.124356 | 1 | 1.123117 |
| 1 | 1.465649 | -0.225776 | 1.239872 | -1.424748 | -0.544383 | 0.110923 | -1.150994 | 0.375698 | -0.600639 | -0.291694 | ... | -1.057711 | 0.822545 | -1.220844 | 0.208864 | -1.959670 | -1.328186 | 0.196861 | -11.263144 | 0 | 2.052266 |
| 2 | 0.738467 | 0.171368 | 0.909835 | -0.301104 | -1.478522 | -0.719844 | -0.460639 | 1.057122 | 0.343618 | -1.763040 | ... | 0.611676 | 1.031000 | 0.931280 | -0.839218 | -0.309212 | 0.331263 | 0.975545 | 0.269378 | 0 | 1.520964 |
| 3 | -0.479174 | -0.185659 | 0.000000 | -1.196207 | 0.812526 | 1.356240 | -0.072010 | 1.003533 | 0.361636 | -0.645120 | ... | 1.564644 | -2.619745 | 0.821903 | 0.087047 | -0.299007 | 0.091761 | -1.987569 | -0.976893 | 0 | 0.125446 |
| 4 | -0.219672 | 0.357113 | 0.137441 | -0.518270 | -0.808494 | -0.501757 | 0.915402 | 0.328751 | -0.529760 | 0.513267 | ... | -0.327662 | -0.392108 | -1.463515 | 0.296120 | 0.261055 | 0.005113 | -0.234587 | -1.949163 | 1 | 0.667889 |
5 rows × 23 columns
[5]:
# Look at the conversion rate and sample size in each group
df.pivot_table(values='outcome',
index='treatment',
aggfunc=[np.mean, np.size],
margins=True)
[5]:
| mean | size | |
|---|---|---|
| outcome | outcome | |
| treatment | ||
| 0 | 0.736125 | 7502 |
| 1 | 1.543688 | 7498 |
| All | 1.139799 | 15000 |
[6]:
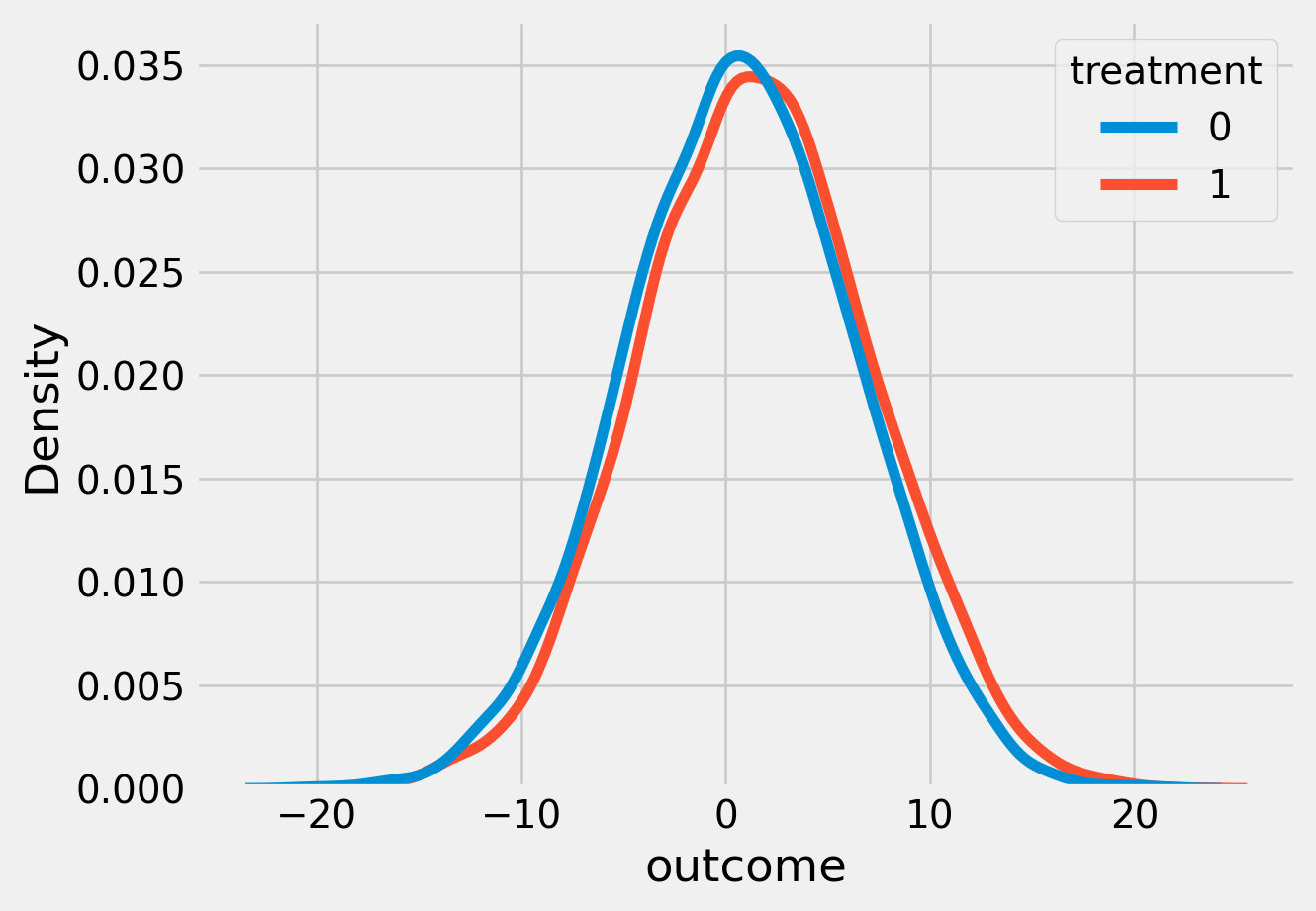
sns.kdeplot(data=df, x='outcome', hue='treatment')
plt.show()
[7]:
# Split data to training and testing samples for model validation (next section)
df_train, df_test = train_test_split(df, test_size=0.2, random_state=11101)
n_test = df_test.shape[0]
n_train = df_train.shape[0]
[8]:
# Table to gather estimated ITEs by models
df_result = pd.DataFrame({
'outcome': df_test['outcome'],
'is_treated': df_test['treatment'],
'treatment_effect': df_test['treatment_effect']
})
CausalTreeRegressor¶
Available criteria for causal trees:
standard_mse: scikit-learn MSE where node values store \(E_{node_i}(X|T=1)-E_{node_i}(X|T=0)\), treatment effects.
causal_mse: The criteria reward a partition for finding strong heterogeneity in treatment effects and penalize a partition that creates variance in leaf estimates. https://www.pnas.org/doi/10.1073/pnas.1510489113
[9]:
ctrees = {
'ctree_mse': {
'params':
dict(criterion='standard_mse',
control_name=0,
min_impurity_decrease=0,
min_samples_leaf=400,
groups_penalty=0.,
groups_cnt=True),
},
'ctree_cmse': {
'params':
dict(
criterion='causal_mse',
control_name=0,
min_samples_leaf=400,
groups_penalty=0.,
groups_cnt=True,
),
},
'ctree_cmse_p=0.1': {
'params':
dict(
criterion='causal_mse',
control_name=0,
min_samples_leaf=400,
groups_penalty=0.1,
groups_cnt=True,
),
},
'ctree_cmse_p=0.25': {
'params':
dict(
criterion='causal_mse',
control_name=0,
min_samples_leaf=400,
groups_penalty=0.25,
groups_cnt=True,
),
},
'ctree_cmse_p=0.5': {
'params':
dict(
criterion='causal_mse',
control_name=0,
min_samples_leaf=400,
groups_penalty=0.5,
groups_cnt=True,
),
},
'ctree_ttest': {
'params':
dict(criterion='t_test',
control_name=0,
min_samples_leaf=400,
groups_penalty=0.,
groups_cnt=True),
},
}
[10]:
# Model treatment effect
for ctree_name, ctree_info in ctrees.items():
print(f"Fitting: {ctree_name}")
ctree = CausalTreeRegressor(**ctree_info['params'])
ctree.fit(X=df_train[feature_names].values,
treatment=df_train['treatment'].values,
y=df_train['outcome'].values)
ctrees[ctree_name].update({'model': ctree})
df_result[ctree_name] = ctree.predict(df_test[feature_names].values)
Fitting: ctree_mse
Fitting: ctree_cmse
Fitting: ctree_cmse_p=0.1
Fitting: ctree_cmse_p=0.25
Fitting: ctree_cmse_p=0.5
Fitting: ctree_ttest
[11]:
df_result.head()
[11]:
| outcome | is_treated | treatment_effect | ctree_mse | ctree_cmse | ctree_cmse_p=0.1 | ctree_cmse_p=0.25 | ctree_cmse_p=0.5 | ctree_ttest | |
|---|---|---|---|---|---|---|---|---|---|
| 625 | 3.519424 | 1 | 0.819201 | 0.110685 | 0.895132 | -1.104810 | 1.166407 | 1.166407 | 0.895132 |
| 5717 | -1.175555 | 0 | 1.131599 | 0.183293 | 3.286218 | 2.071375 | 0.794040 | 0.794040 | 2.096099 |
| 14801 | 4.361167 | 0 | 1.969727 | 0.834163 | 3.329034 | 3.497691 | 3.097263 | 3.097263 | 2.077012 |
| 13605 | 4.523891 | 0 | 0.884079 | 0.183293 | -0.727168 | -2.025098 | -0.902955 | -0.902955 | -0.727168 |
| 4208 | -6.077212 | 0 | 1.179124 | 0.183293 | 3.329034 | 3.497691 | 1.916049 | 1.916049 | 1.257711 |
[12]:
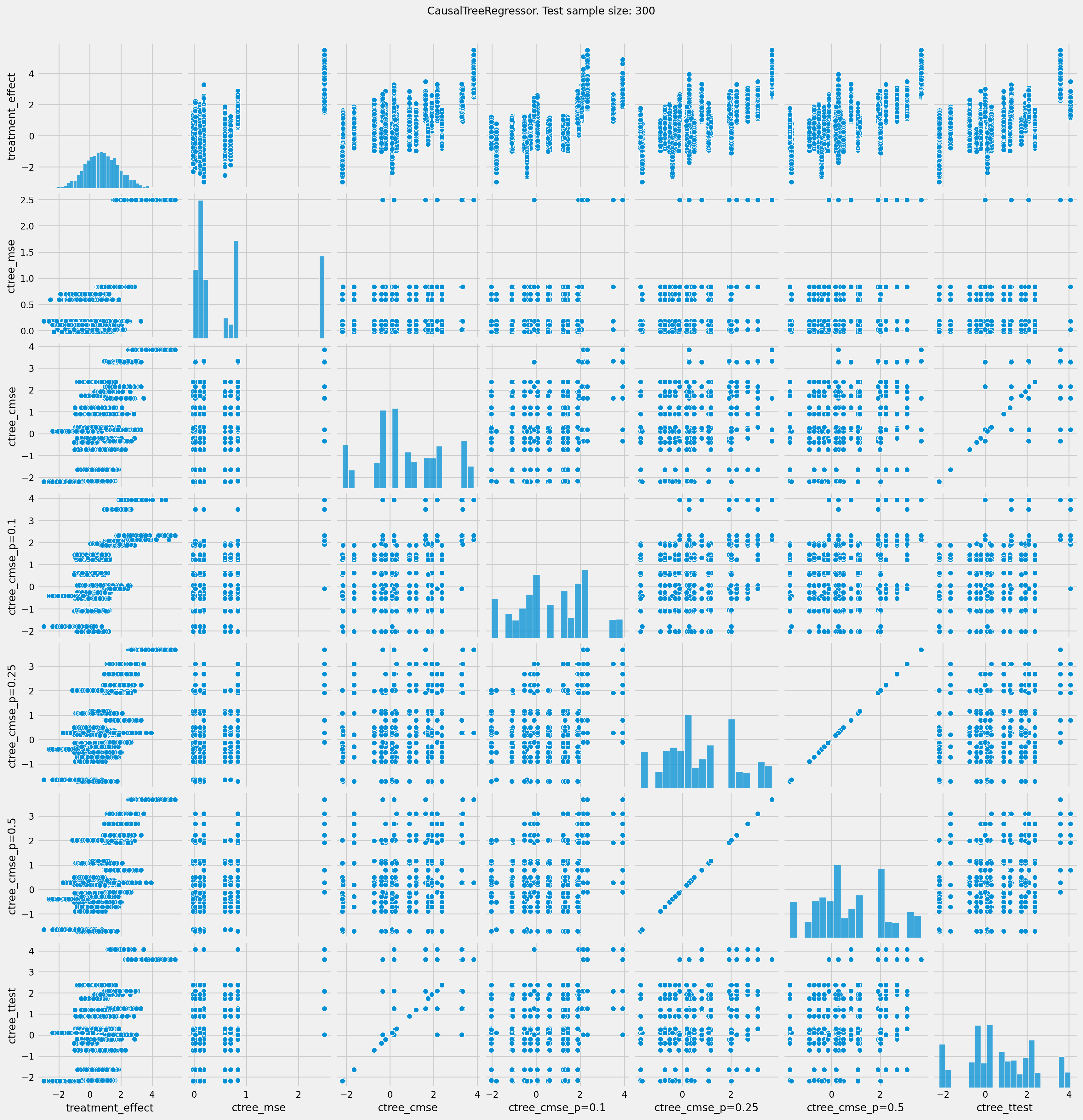
# See treatment effect estimation with CausalTreeRegressor vs true treatment effect
n_obs = 300
indxs = df_result.index.values
np.random.shuffle(indxs)
indxs = indxs[:n_obs]
plt.rcParams.update({'font.size': 10})
pairplot = sns.pairplot(df_result[['treatment_effect', *list(ctrees)]])
pairplot.fig.suptitle(f"CausalTreeRegressor. Test sample size: {n_obs}" , y=1.02)
plt.show()
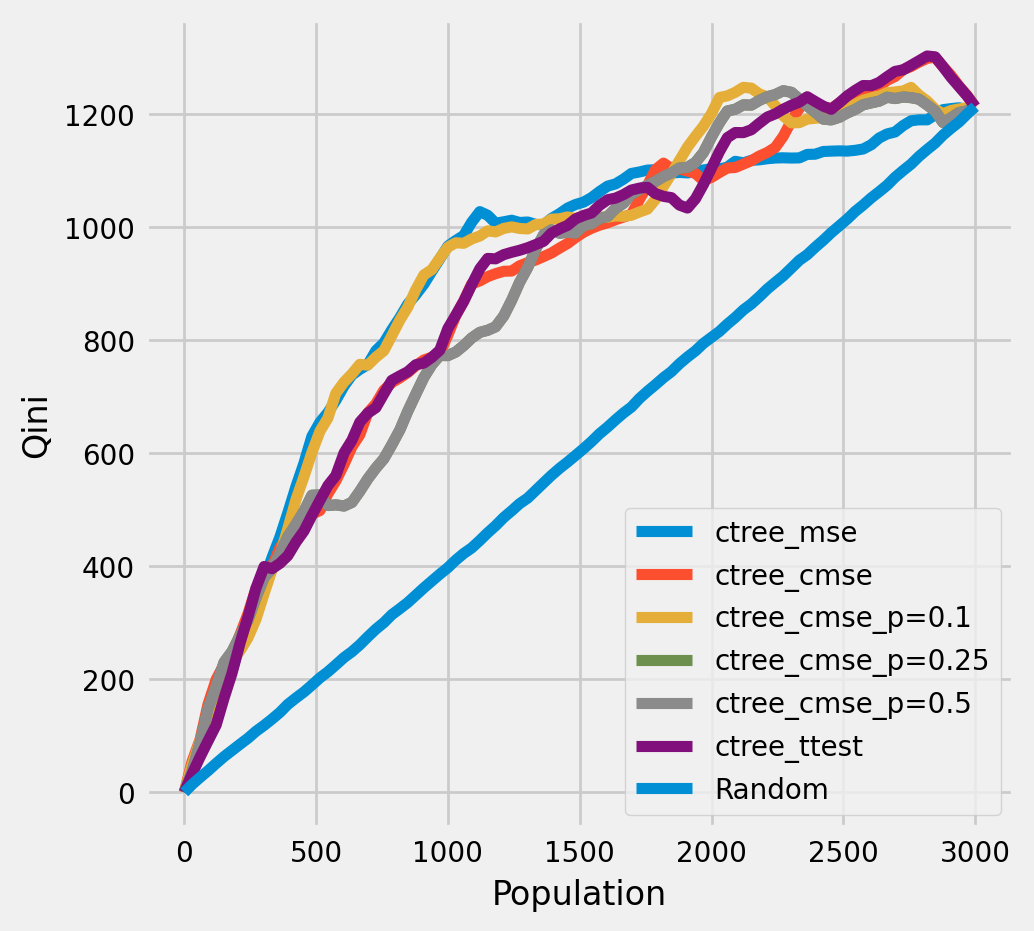
Plot the Qini chart¶
[13]:
plot_qini(df_result,
outcome_col='outcome',
treatment_col='is_treated',
treatment_effect_col='treatment_effect',
figsize=(5,5)
)
[14]:
df_qini = qini_score(df_result,
outcome_col='outcome',
treatment_col='is_treated',
treatment_effect_col='treatment_effect')
df_qini.sort_values(ascending=False)
[14]:
ctree_cmse_p=0.1 0.273112
ctree_mse 0.260141
ctree_ttest 0.247668
ctree_cmse 0.242333
ctree_cmse_p=0.25 0.231232
ctree_cmse_p=0.5 0.231232
Random 0.000000
dtype: float64
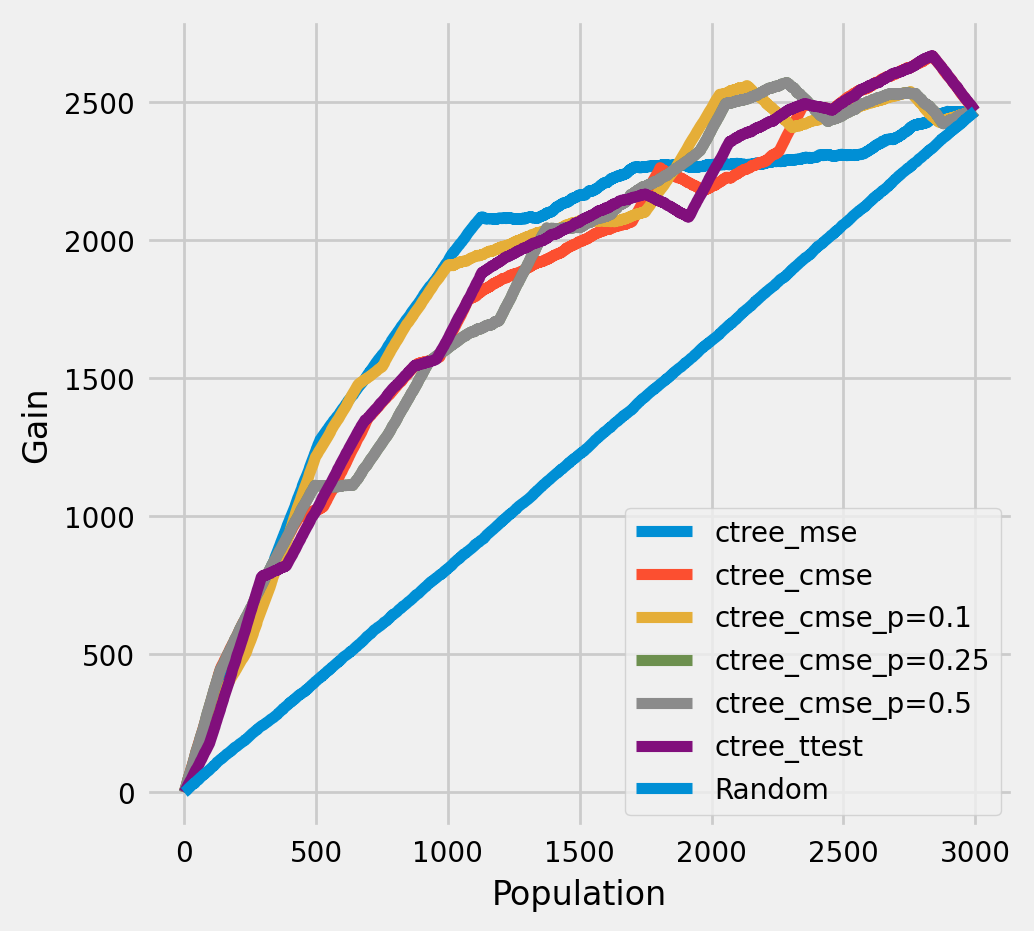
The cumulative gain of the true treatment effect in each population¶
[15]:
plot_gain(df_result,
outcome_col='outcome',
treatment_col='is_treated',
treatment_effect_col='treatment_effect',
n = n_test,
figsize=(5,5)
)
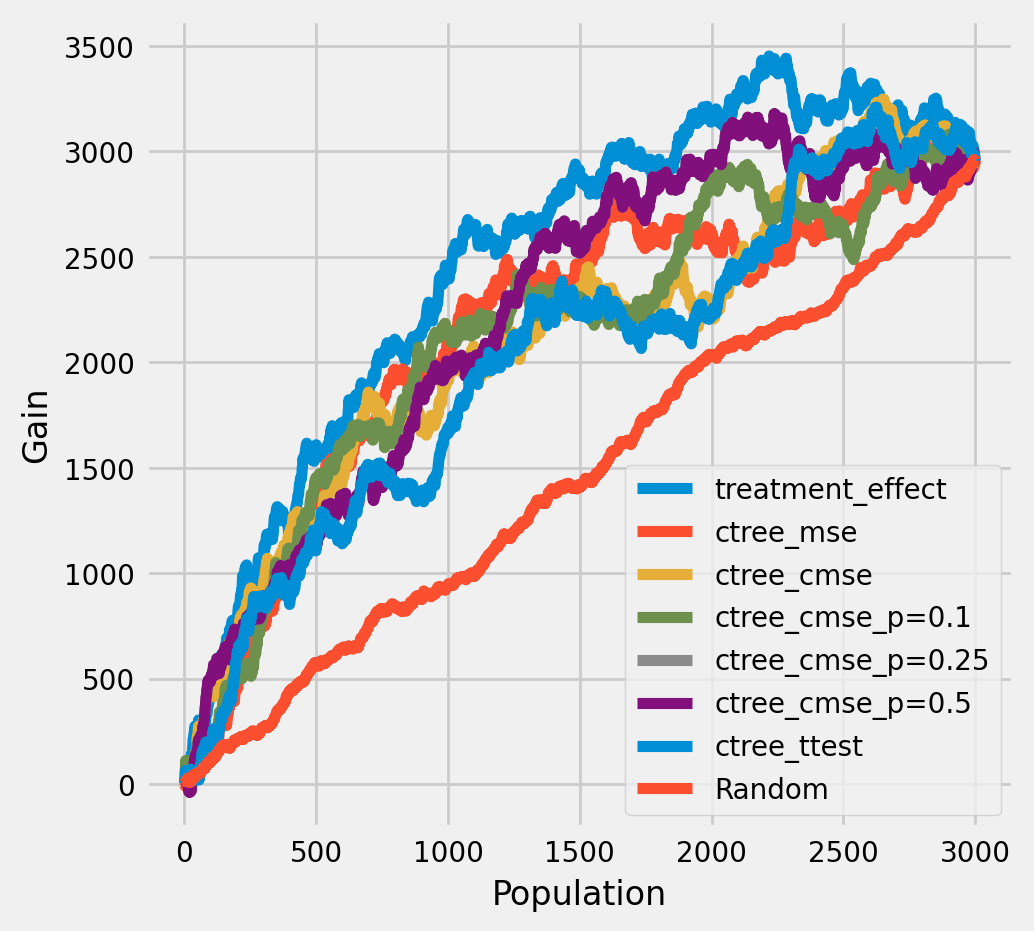
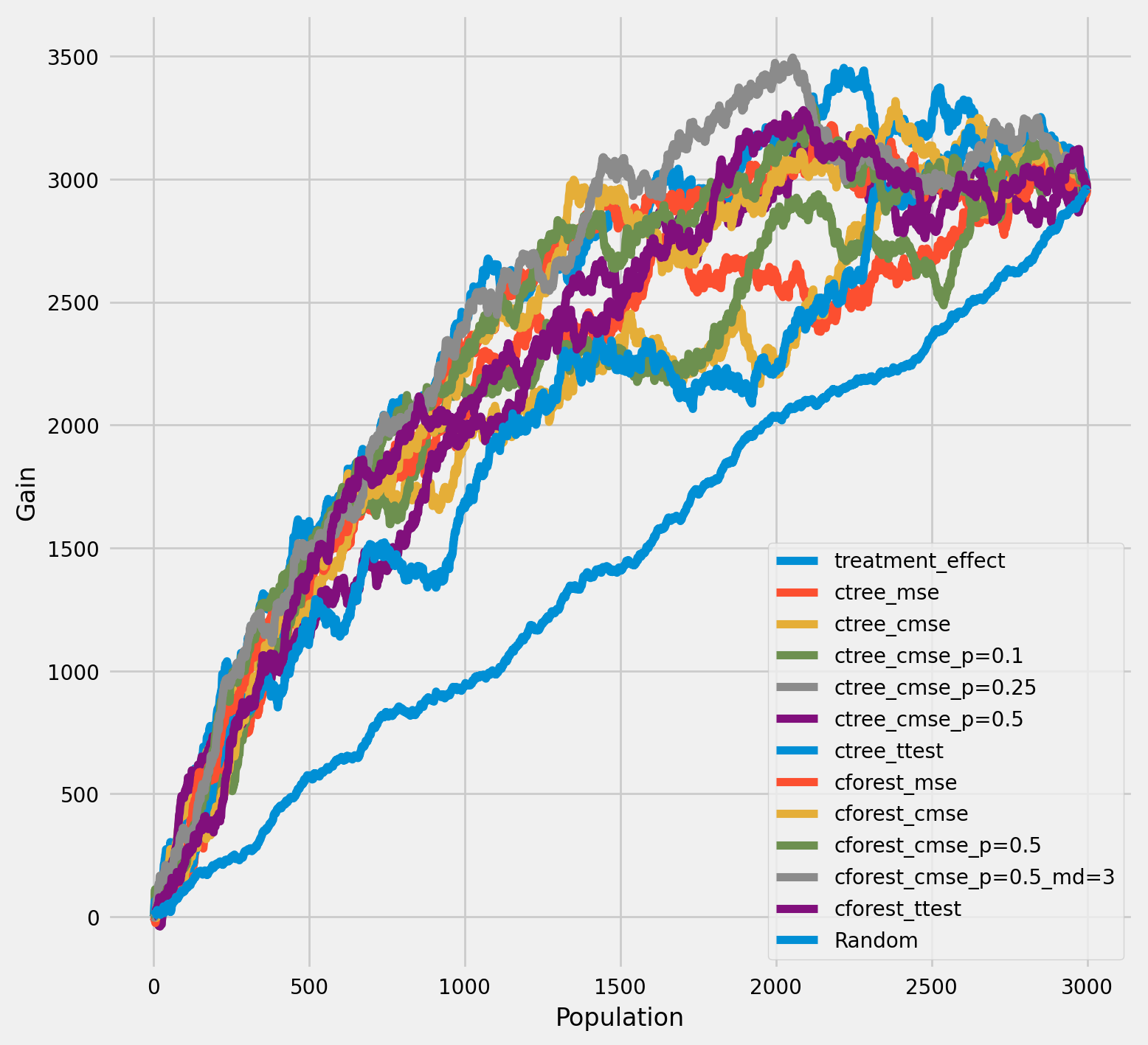
The cumulative difference between the mean outcomes of the treatment and control groups in each population¶
[16]:
plot_gain(df_result,
outcome_col='outcome',
treatment_col='is_treated',
n = n_test,
figsize=(5,5)
)
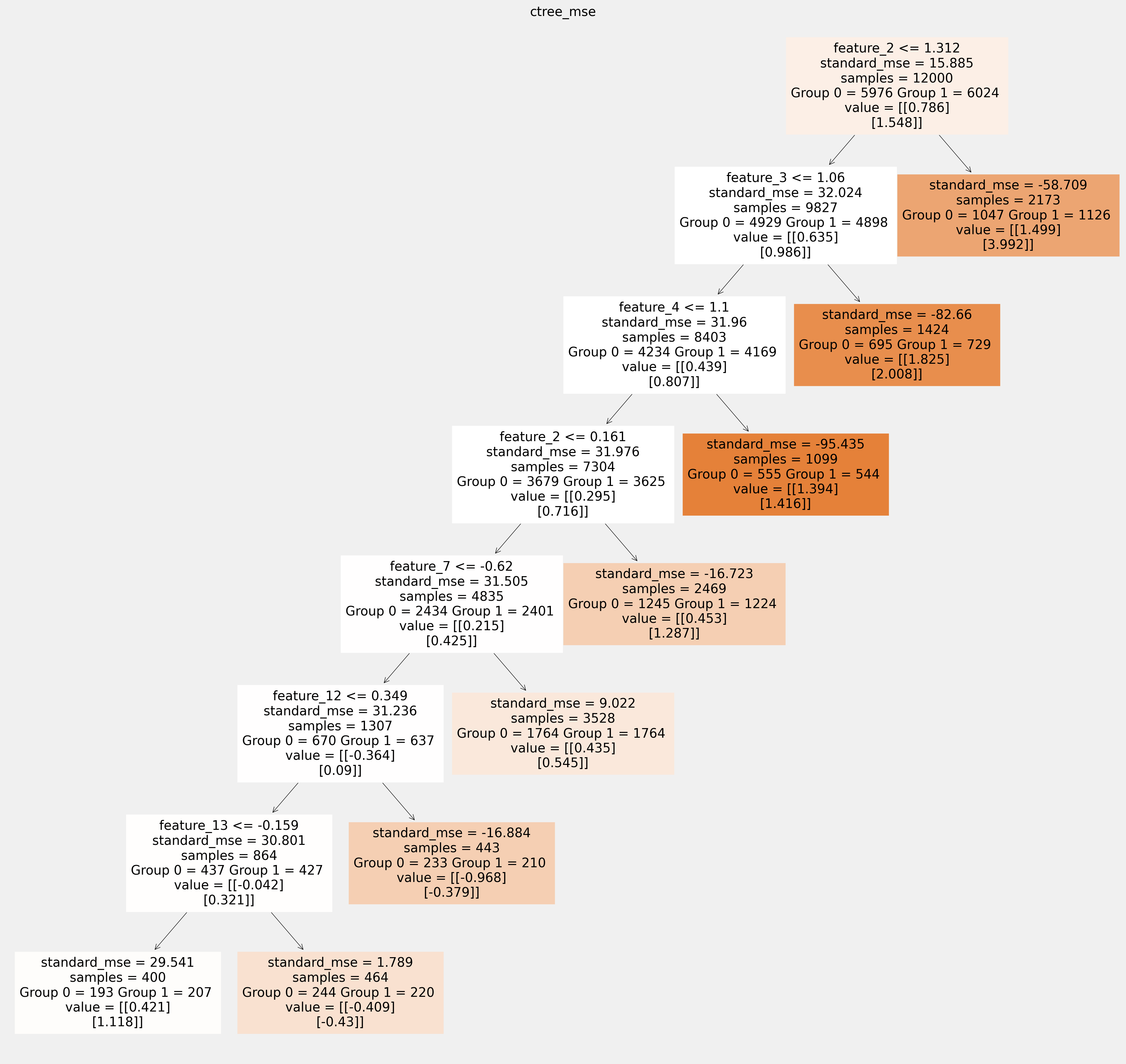
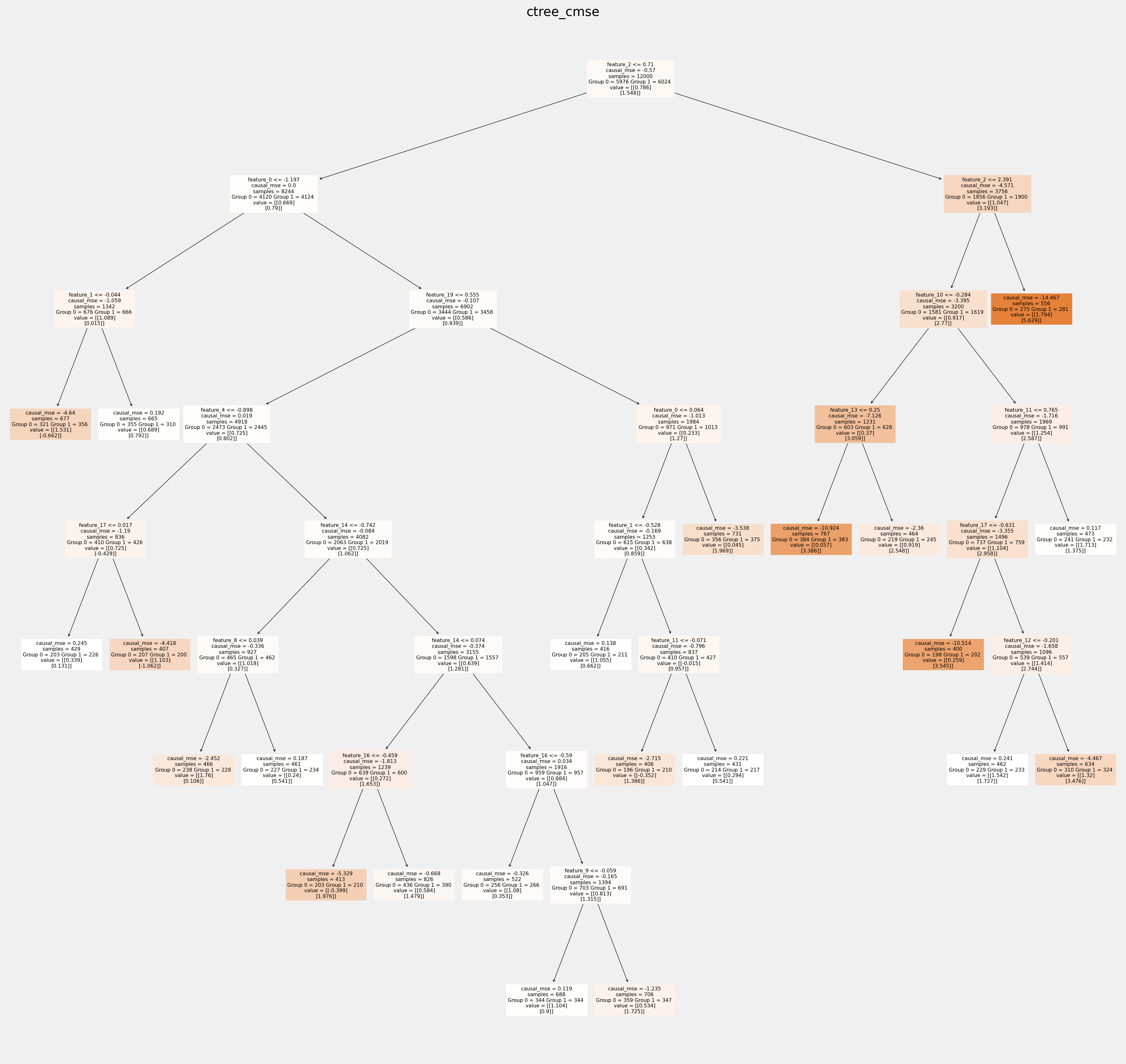
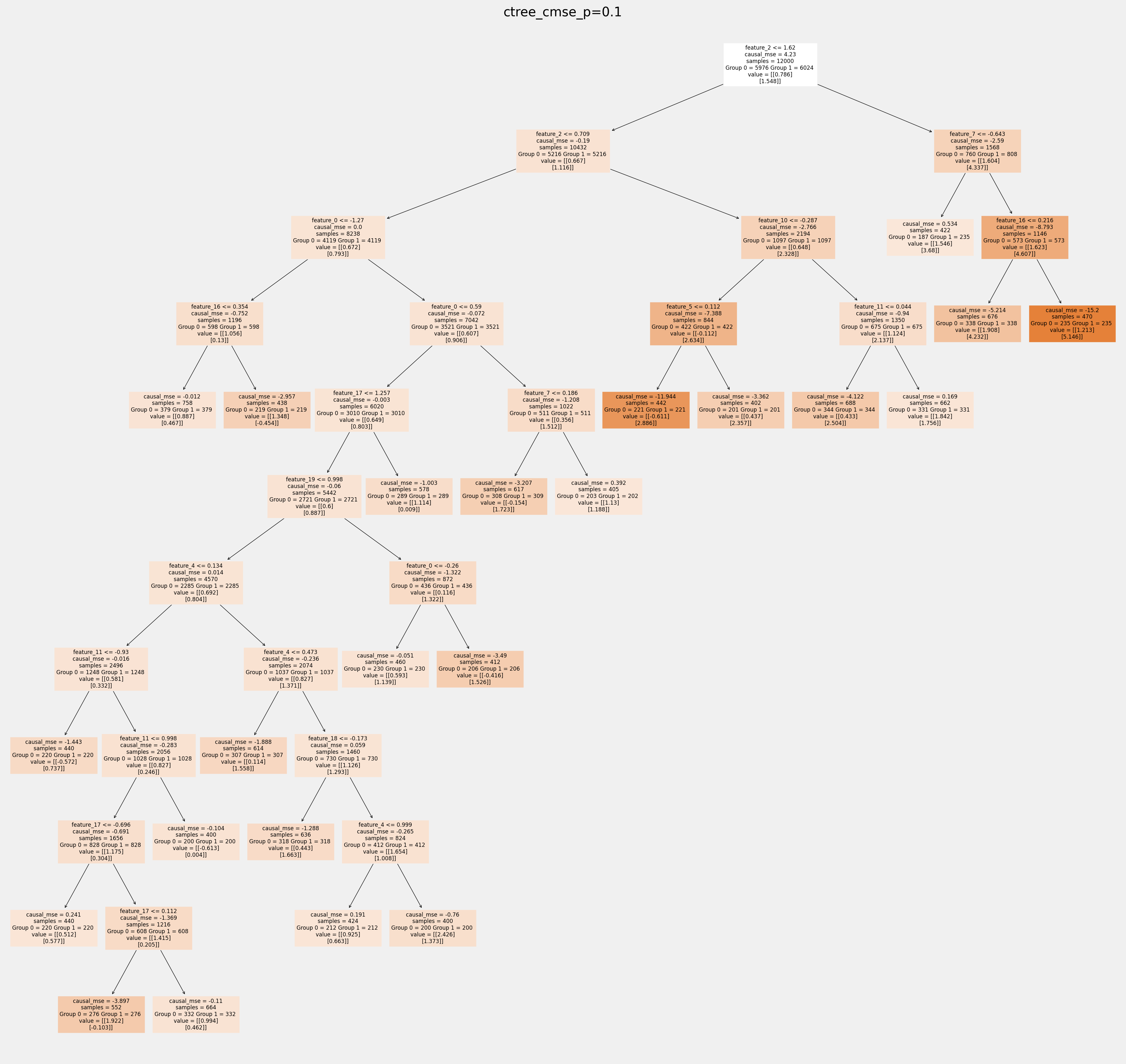
Plot trees with sklearn function and save as vector graphics¶
[17]:
for ctree_name, ctree_info in ctrees.items():
plt.figure(figsize=(20,20))
plot_causal_tree(ctree_info['model'],
feature_names = feature_names,
filled=True,
impurity=True,
proportion=False,
)
plt.title(ctree_name)
plt.savefig(f'{ctree_name}.svg')
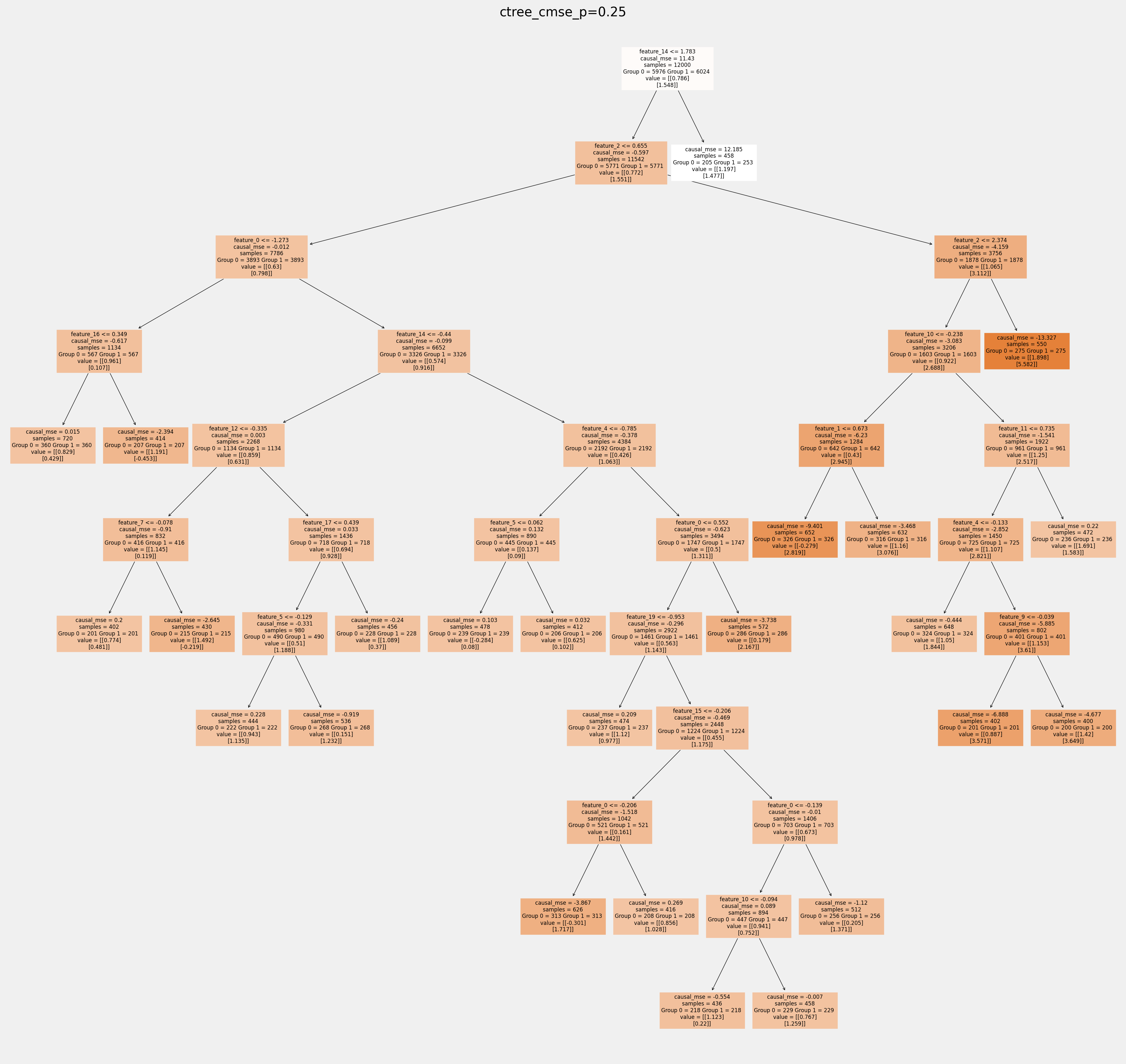
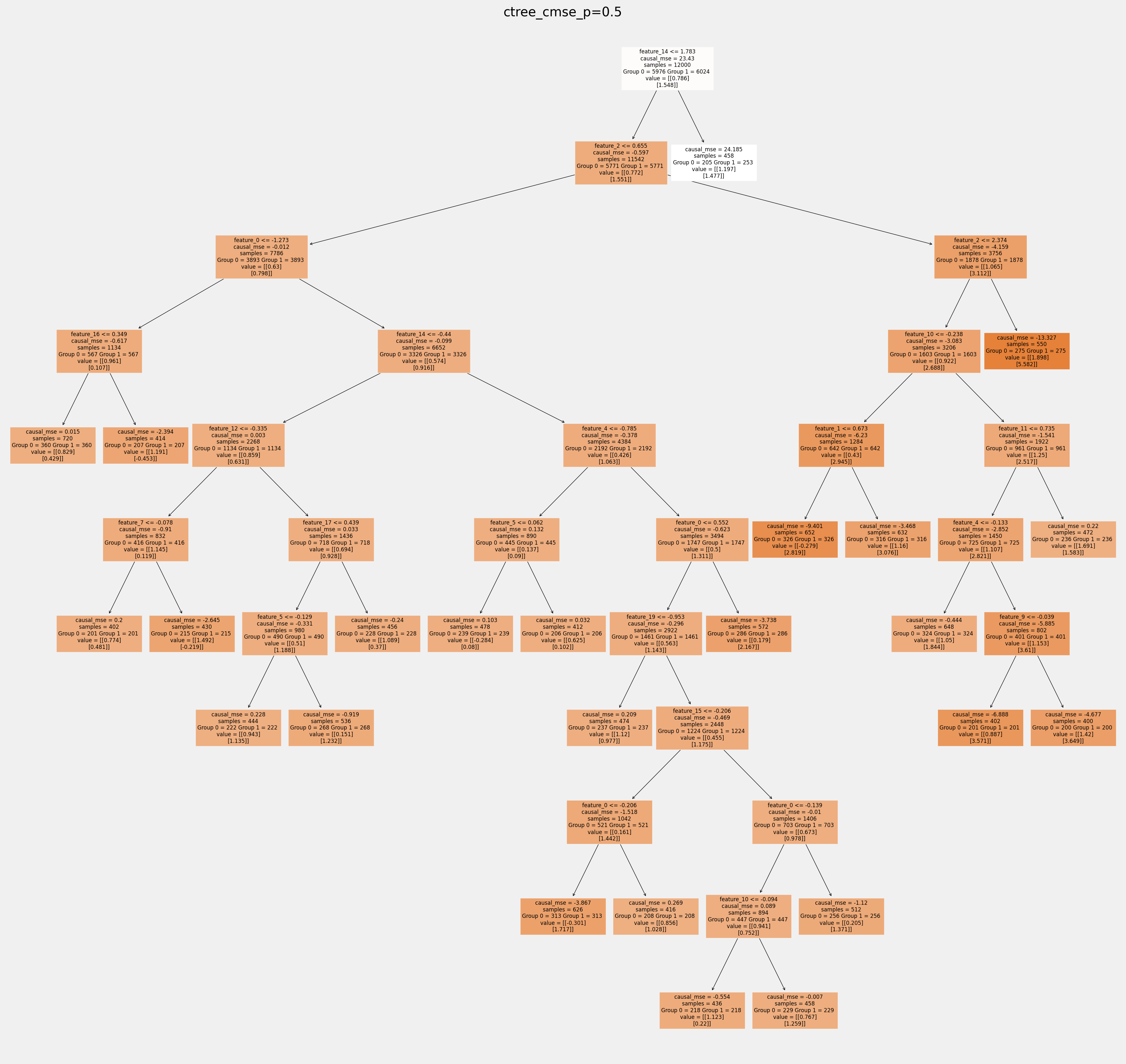
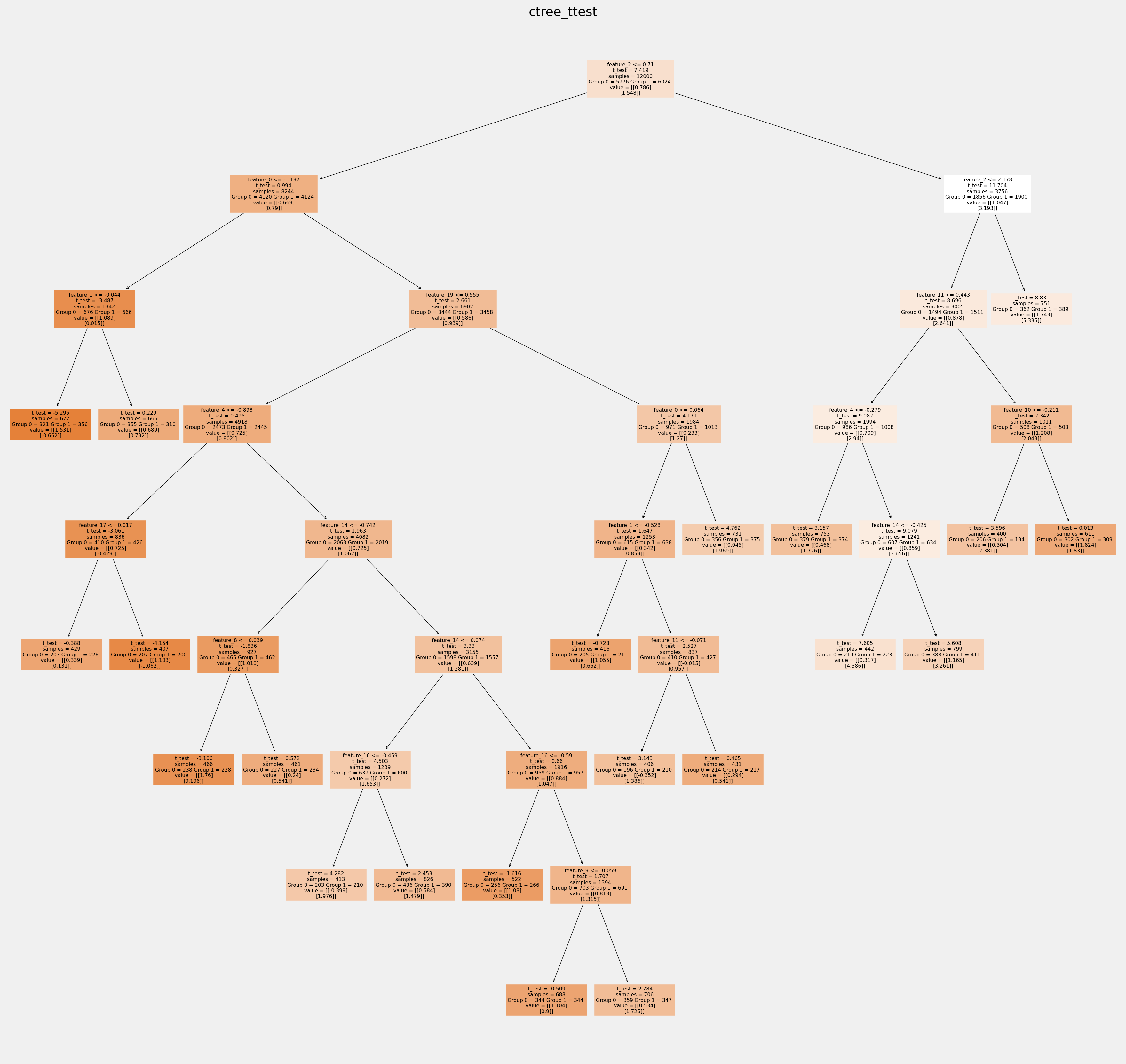
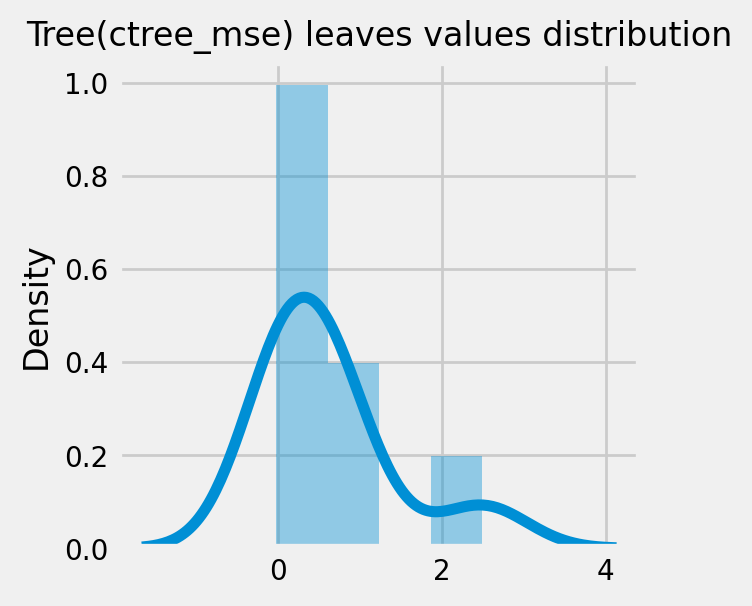
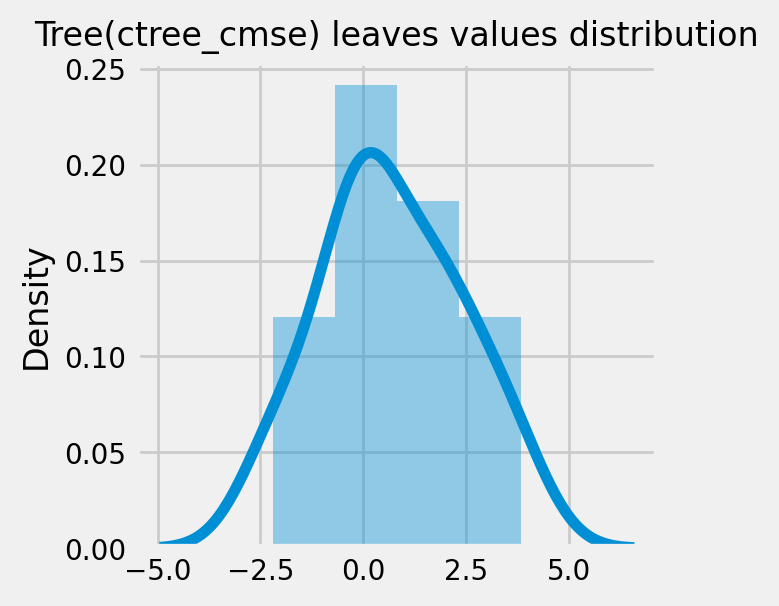
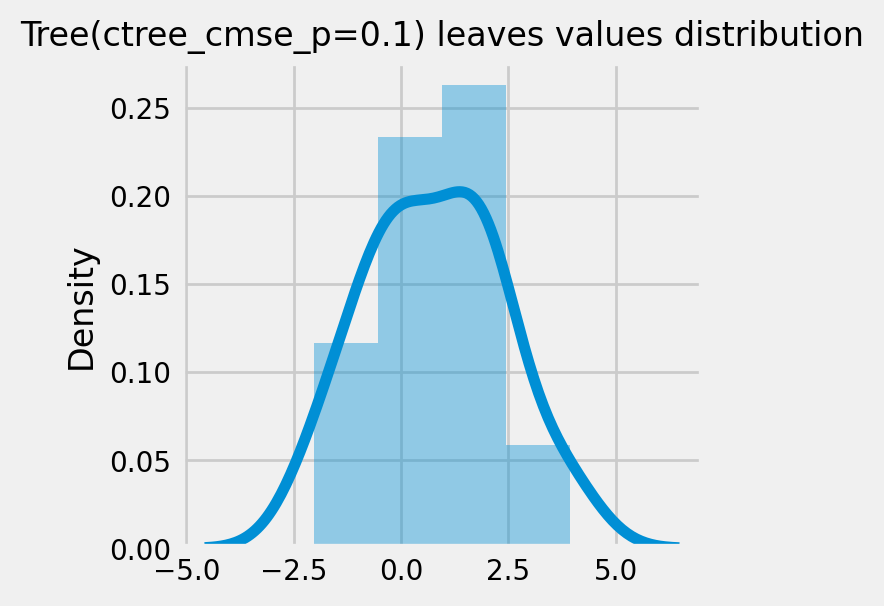
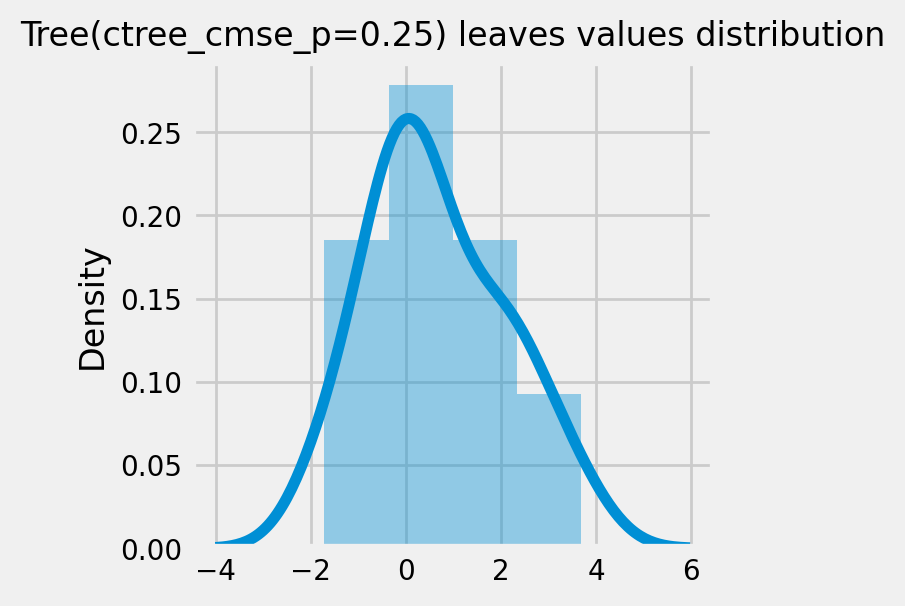
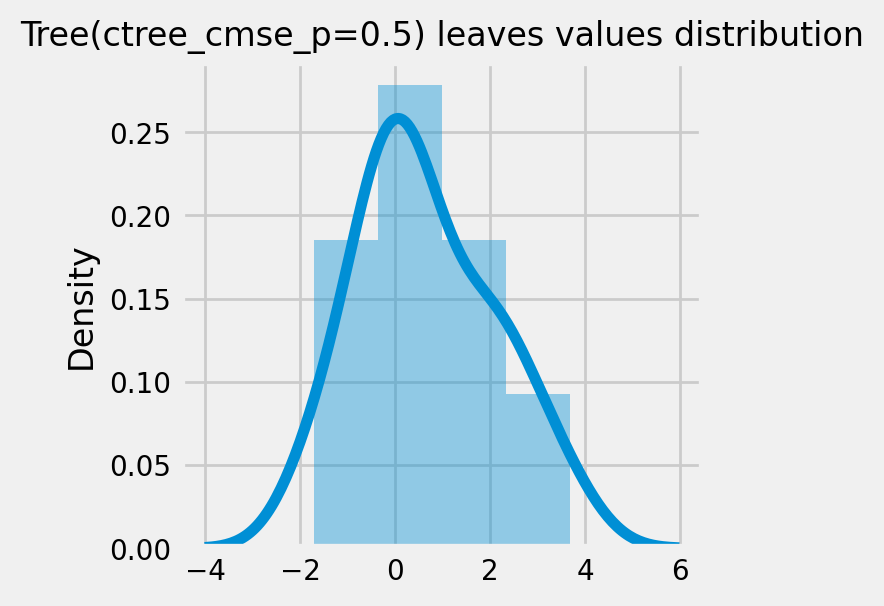
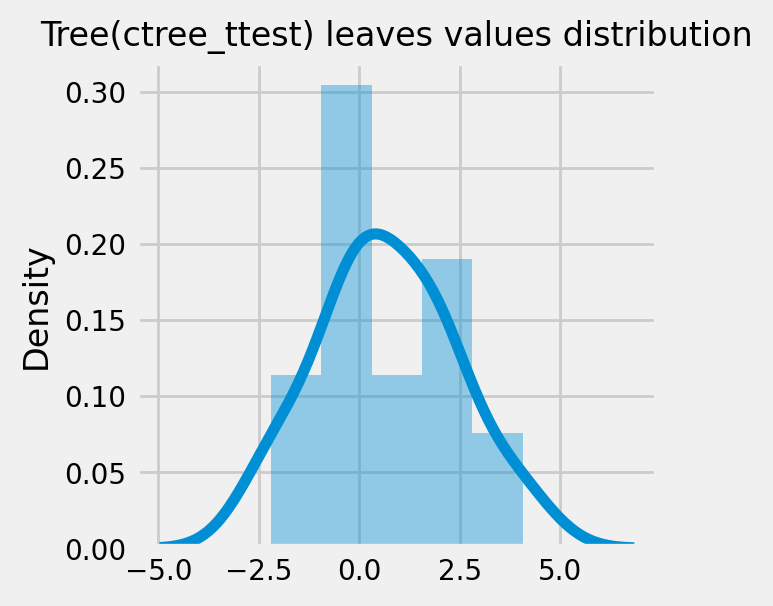
How values in leaves of the fitted trees differ from each other:¶
[18]:
for ctree_name, ctree_info in ctrees.items():
plot_dist_tree_leaves_values(ctree_info['model'],
figsize=(3,3),
title=f'Tree({ctree_name}) leaves values distribution')
CausalRandomForestRegressor¶
[19]:
cforests = {
'cforest_mse': {
'params':
dict(criterion='standard_mse',
control_name=0,
min_impurity_decrease=0,
min_samples_leaf=400,
groups_penalty=0.,
groups_cnt=True),
},
'cforest_cmse': {
'params':
dict(
criterion='causal_mse',
control_name=0,
min_samples_leaf=400,
groups_penalty=0.,
groups_cnt=True
),
},
'cforest_cmse_p=0.5': {
'params':
dict(
criterion='causal_mse',
control_name=0,
min_samples_leaf=400,
groups_penalty=0.5,
groups_cnt=True,
),
},
'cforest_cmse_p=0.5_md=3': {
'params':
dict(
criterion='causal_mse',
control_name=0,
max_depth=3,
min_samples_leaf=400,
groups_penalty=0.5,
groups_cnt=True,
),
},
'cforest_ttest': {
'params':
dict(criterion='t_test',
control_name=0,
min_samples_leaf=400,
groups_penalty=0.,
groups_cnt=True),
},
}
[20]:
# Model treatment effect
for cforest_name, cforest_info in cforests.items():
print(f"Fitting: {cforest_name}")
cforest = CausalRandomForestRegressor(**cforest_info['params'])
cforest.fit(X=df_train[feature_names].values,
treatment=df_train['treatment'].values,
y=df_train['outcome'].values)
cforests[cforest_name].update({'model': cforest})
df_result[cforest_name] = cforest.predict(df_test[feature_names].values)
Fitting: cforest_mse
Fitting: cforest_cmse
Fitting: cforest_cmse_p=0.5
Fitting: cforest_cmse_p=0.5_md=3
Fitting: cforest_ttest
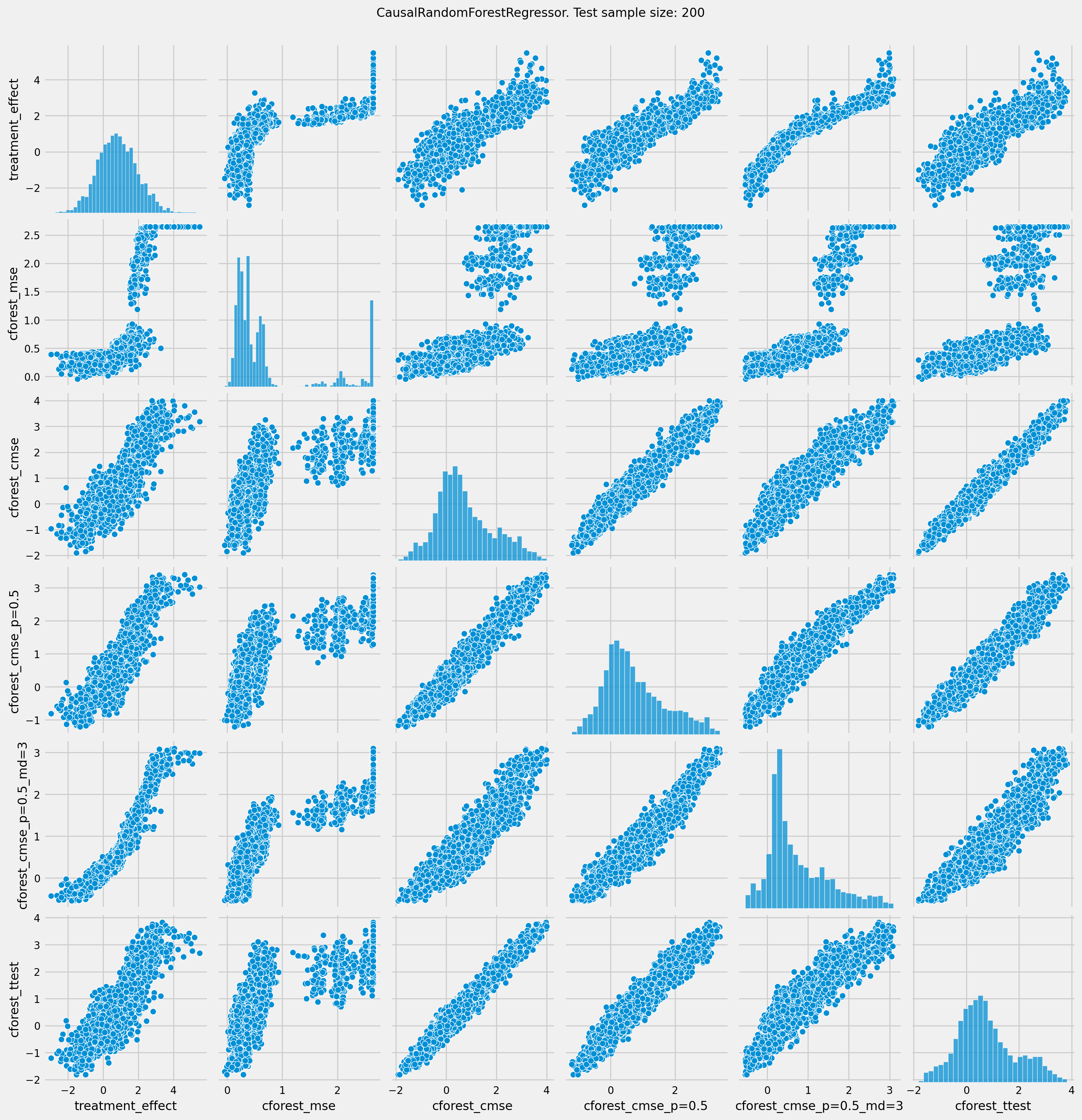
[21]:
# See treatment effect estimation with CausalRandomForestRegressor vs true treatment effect
n_obs = 200
indxs = df_result.index.values
np.random.shuffle(indxs)
indxs = indxs[:n_obs]
plt.rcParams.update({'font.size': 10})
pairplot = sns.pairplot(df_result[['treatment_effect', *list(cforests)]])
pairplot.fig.suptitle(f"CausalRandomForestRegressor. Test sample size: {n_obs}" , y=1.02)
plt.show()
[22]:
df_qini = qini_score(df_result,
outcome_col='outcome',
treatment_col='is_treated',
treatment_effect_col='treatment_effect')
df_qini.sort_values(ascending=False)
[22]:
cforest_cmse_p=0.5_md=3 0.379598
cforest_cmse_p=0.5 0.344870
cforest_ttest 0.329592
cforest_mse 0.326770
cforest_cmse 0.323620
ctree_cmse_p=0.1 0.273112
ctree_mse 0.260141
ctree_ttest 0.247668
ctree_cmse 0.242333
ctree_cmse_p=0.25 0.231232
ctree_cmse_p=0.5 0.231232
Random 0.000000
dtype: float64
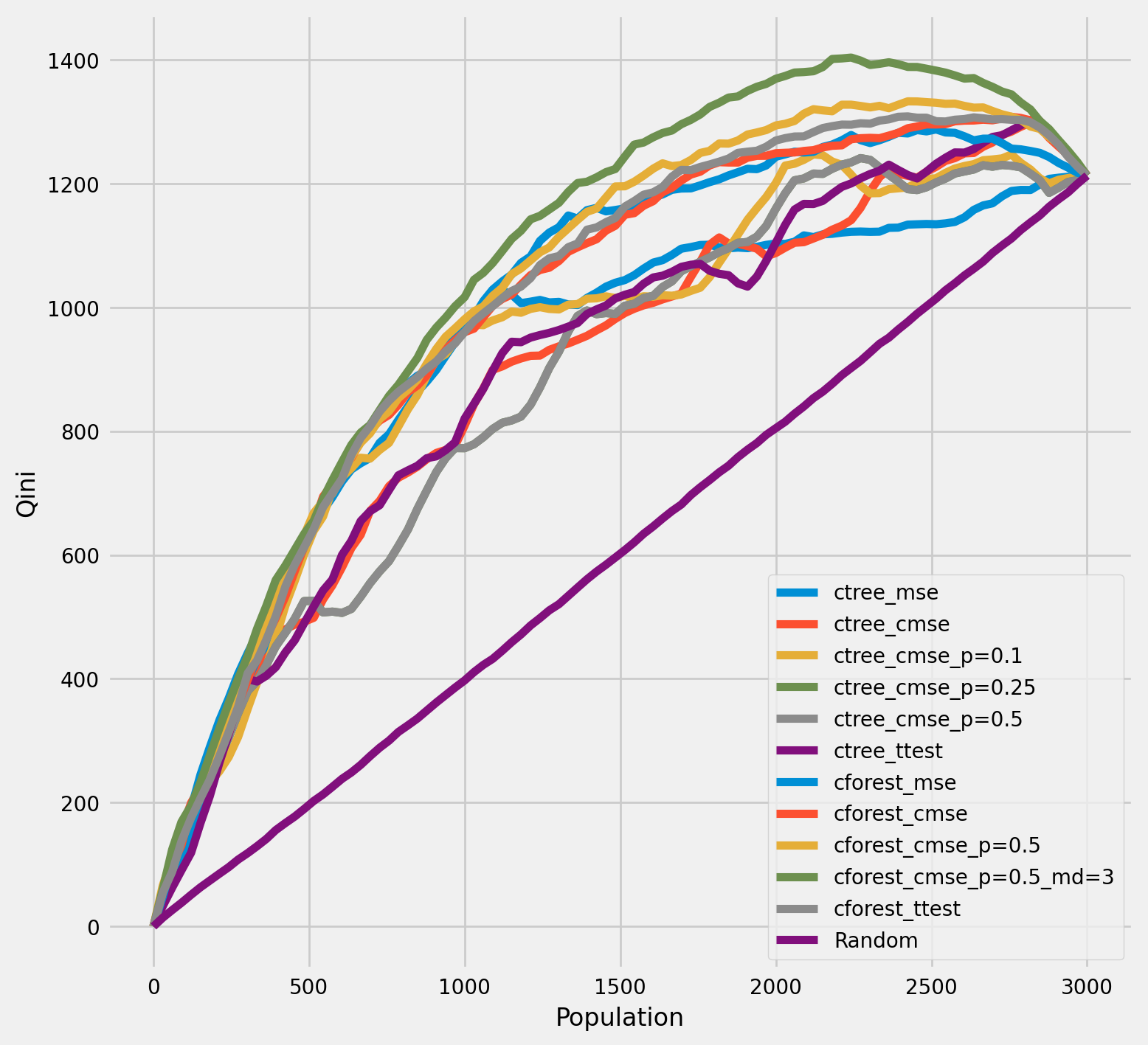
Qini chart¶
[23]:
plot_qini(df_result,
outcome_col='outcome',
treatment_col='is_treated',
treatment_effect_col='treatment_effect',
figsize=(8,8)
)
[24]:
df_qini = qini_score(df_result,
outcome_col='outcome',
treatment_col='is_treated',
treatment_effect_col='treatment_effect')
df_qini.sort_values(ascending=False)
[24]:
cforest_cmse_p=0.5_md=3 0.379598
cforest_cmse_p=0.5 0.344870
cforest_ttest 0.329592
cforest_mse 0.326770
cforest_cmse 0.323620
ctree_cmse_p=0.1 0.273112
ctree_mse 0.260141
ctree_ttest 0.247668
ctree_cmse 0.242333
ctree_cmse_p=0.25 0.231232
ctree_cmse_p=0.5 0.231232
Random 0.000000
dtype: float64
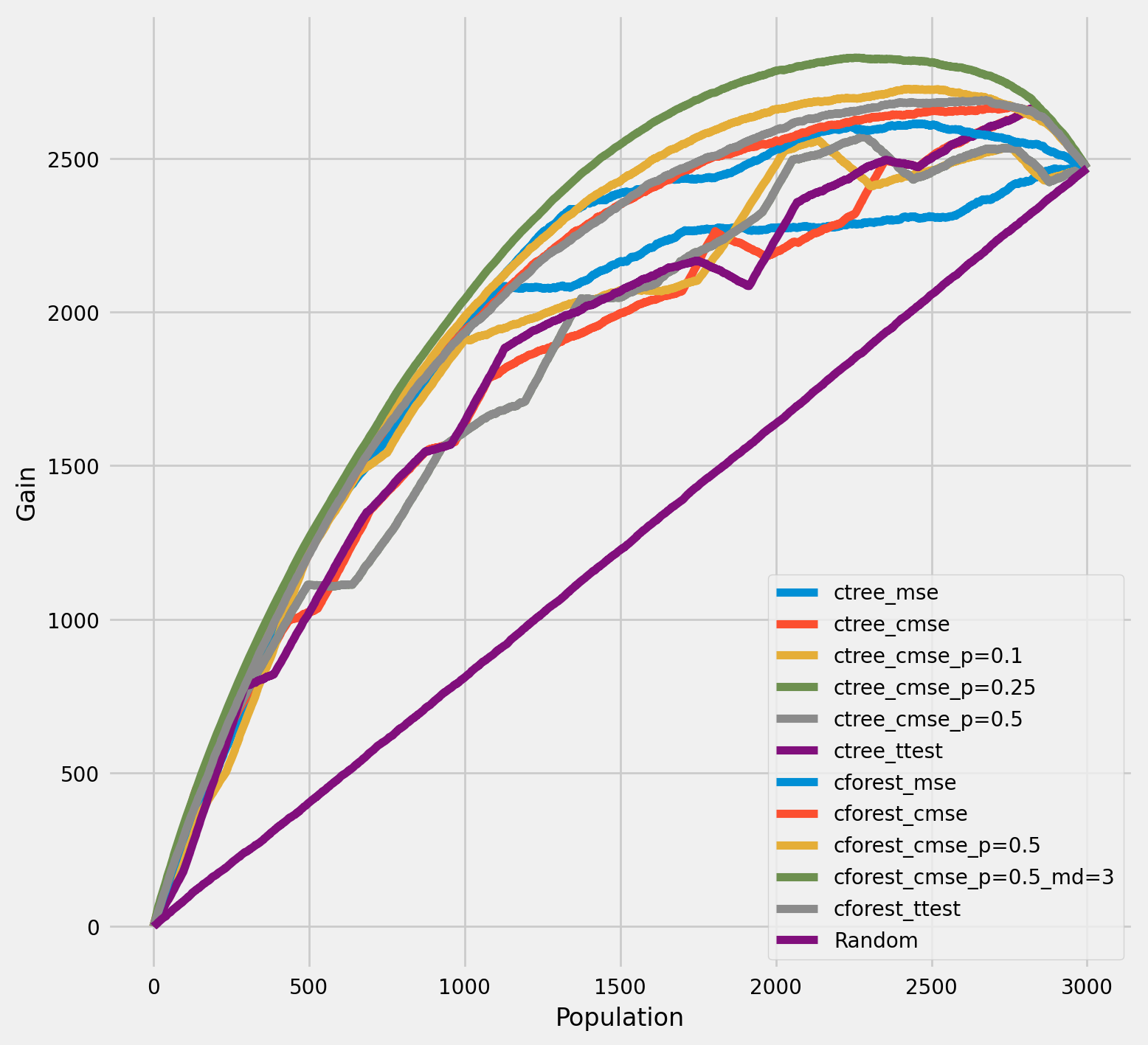
The cumulative gain of the true treatment effect in each population¶
[25]:
plot_gain(df_result,
outcome_col='outcome',
treatment_col='is_treated',
treatment_effect_col='treatment_effect',
n = n_test
)
The cumulative difference between the mean outcomes of the treatment and control groups in each population¶
[26]:
plot_gain(df_result,
outcome_col='outcome',
treatment_col='is_treated',
n = n_test
)
Meta-Learner Algorithms¶
[27]:
X_train = df_train[feature_names].values
X_test = df_test[feature_names].values
# learner - DecisionTreeRegressor
# treatment learner - LinearRegression()
learner_x = BaseXRegressor(learner=DecisionTreeRegressor(),
treatment_effect_learner=LinearRegression())
learner_s = BaseSRegressor(learner=DecisionTreeRegressor())
learner_t = BaseTRegressor(learner=DecisionTreeRegressor(),
treatment_learner=LinearRegression())
learner_dr = BaseDRRegressor(learner=DecisionTreeRegressor(),
treatment_effect_learner=LinearRegression())
learner_x.fit(X=X_train, treatment=df_train['treatment'].values, y=df_train['outcome'].values)
learner_s.fit(X=X_train, treatment=df_train['treatment'].values, y=df_train['outcome'].values)
learner_t.fit(X=X_train, treatment=df_train['treatment'].values, y=df_train['outcome'].values)
learner_dr.fit(X=X_train, treatment=df_train['treatment'].values, y=df_train['outcome'].values)
df_result['learner_x_ite'] = learner_x.predict(X_test)
df_result['learner_s_ite'] = learner_s.predict(X_test)
df_result['learner_t_ite'] = learner_t.predict(X_test)
df_result['learner_dr_ite'] = learner_dr.predict(X_test)
[28]:
cate_dr = learner_dr.predict(X)
cate_x = learner_x.predict(X)
cate_s = learner_s.predict(X)
cate_t = learner_t.predict(X)
cate_ctrees = [info['model'].predict(X) for _, info in ctrees.items()]
cate_cforests = [info['model'].predict(X) for _, info in cforests.items()]
model_cate = [
*cate_ctrees,
*cate_cforests,
cate_x, cate_s, cate_t, cate_dr
]
model_names = [
*list(ctrees), *list(cforests),
'X Learner', 'S Learner', 'T Learner', 'DR Learner']
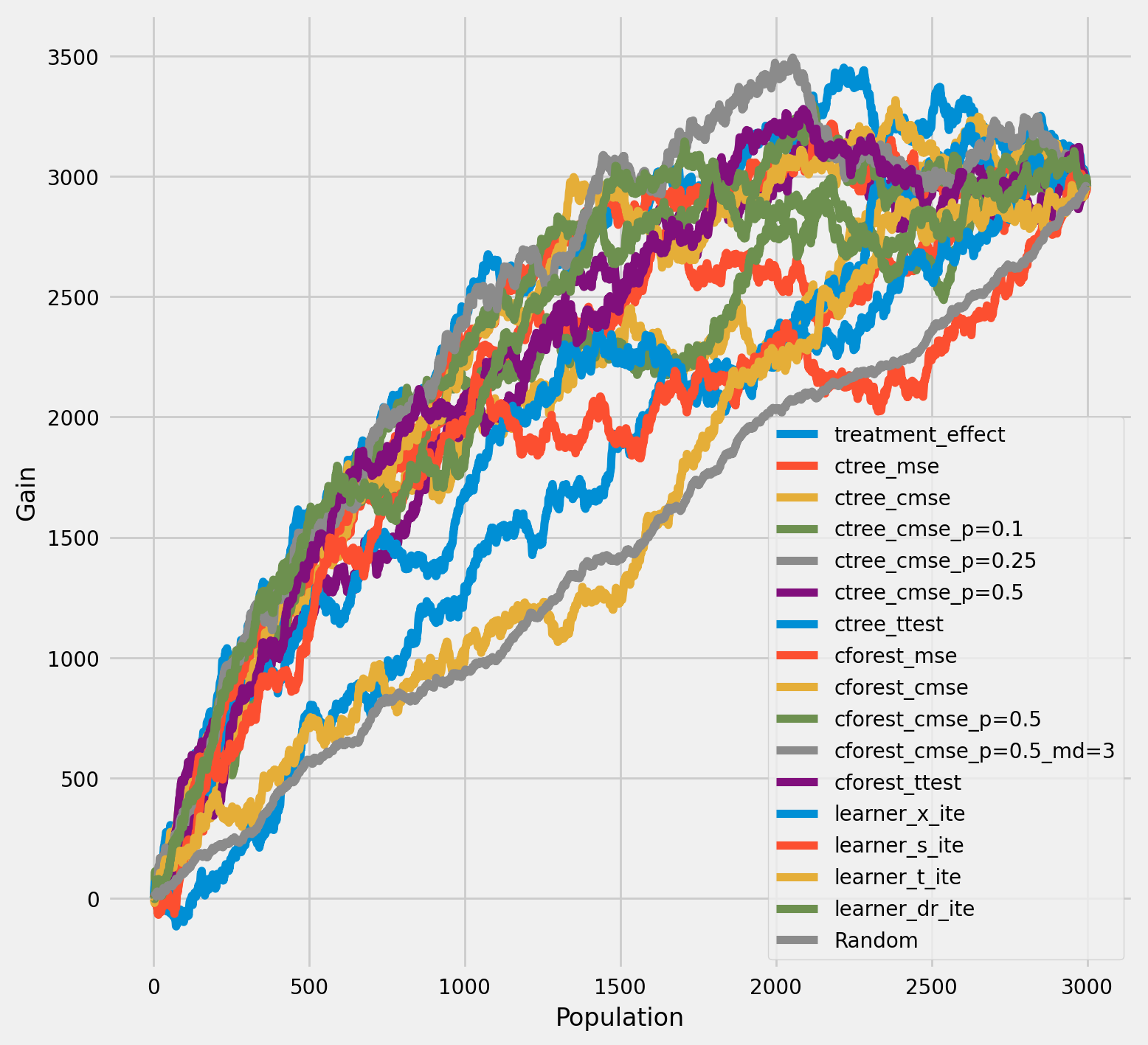
[29]:
plot_gain(df_result,
outcome_col='outcome',
treatment_col='is_treated',
n = n_test
)
[30]:
rows = 2
cols = 7
row_idxs = np.arange(rows)
col_idxs = np.arange(cols)
ax_idxs = np.dstack(np.meshgrid(col_idxs, row_idxs)).reshape(-1, 2)
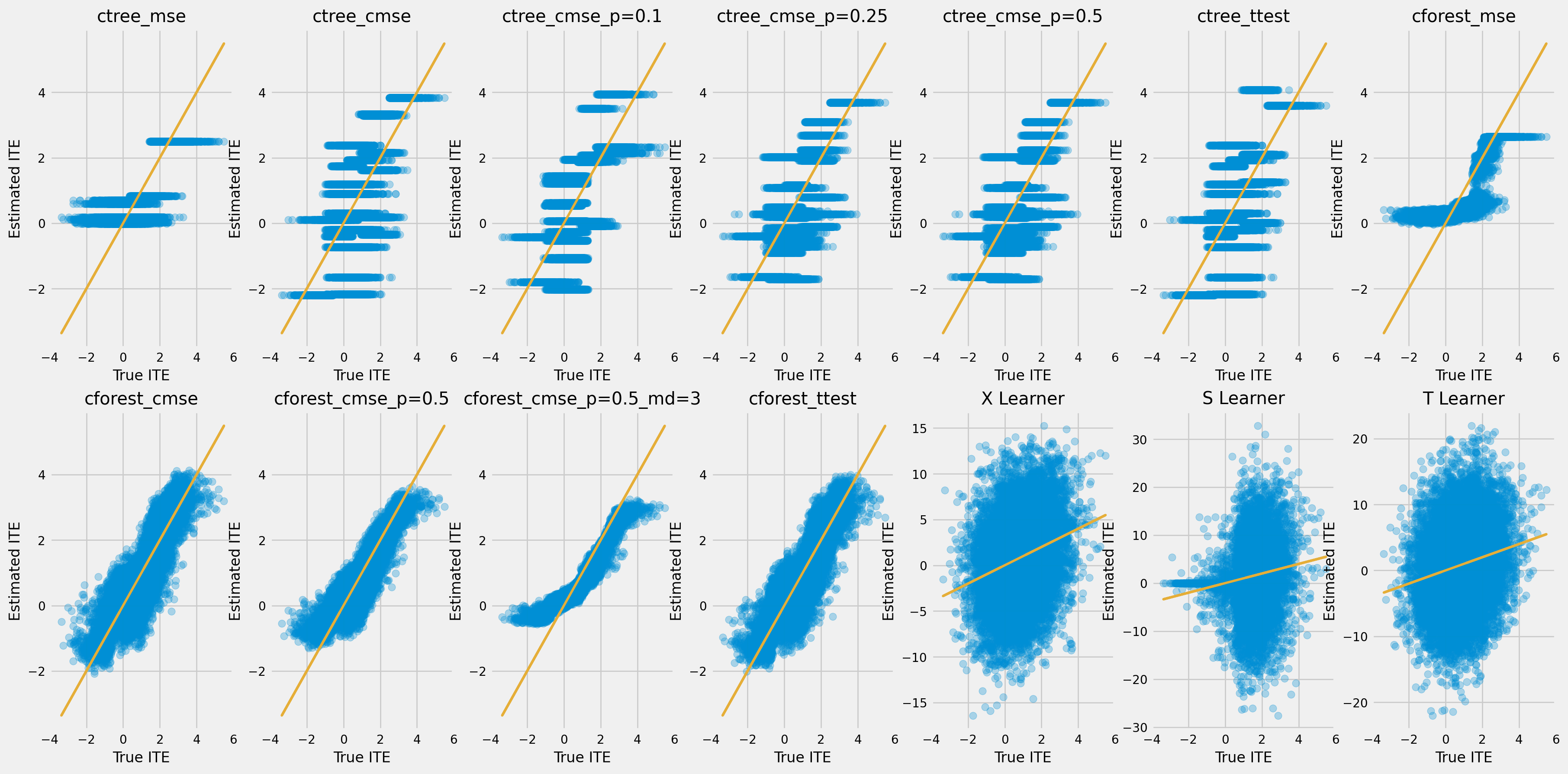
[31]:
fig, ax = plt.subplots(rows, cols, figsize=(20, 10))
plt.rcParams.update({'font.size': 10})
for ax_idx, cate, model_name in zip(ax_idxs, model_cate, model_names):
col_id, row_id = ax_idx
cur_ax = ax[row_id, col_id]
cur_ax.scatter(tau, cate, alpha=0.3)
cur_ax.plot(tau, tau, color='C2', linewidth=2)
cur_ax.set_xlabel('True ITE')
cur_ax.set_ylabel('Estimated ITE')
cur_ax.set_title(model_name)
cur_ax.set_xlim((-4, 6))
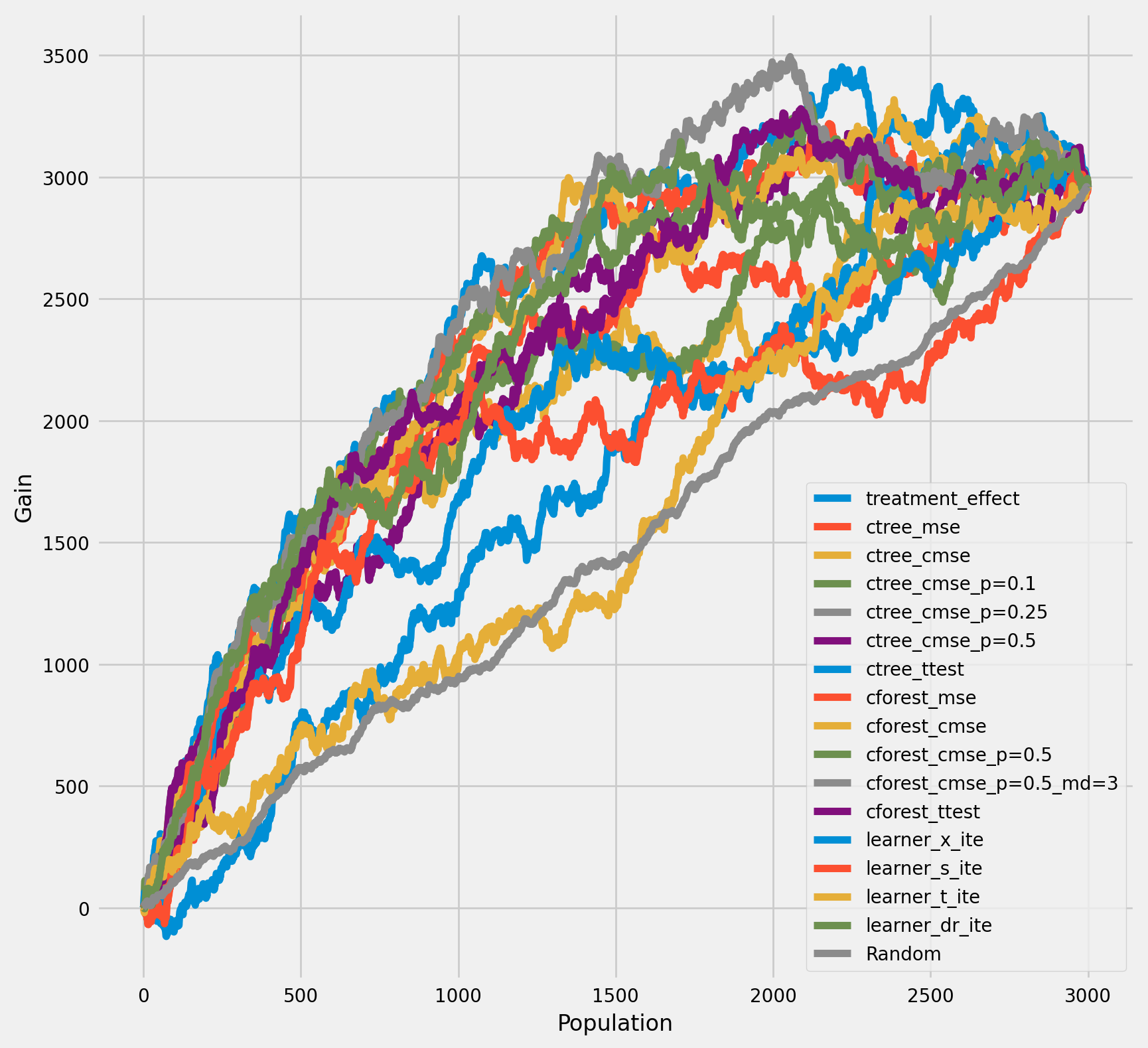
The cumulative difference between the mean outcomes of the treatment and control groups in each population¶
[32]:
plot_gain(df_result,
outcome_col='outcome',
treatment_col='is_treated',
n = n_test,
figsize=(9, 9),
)
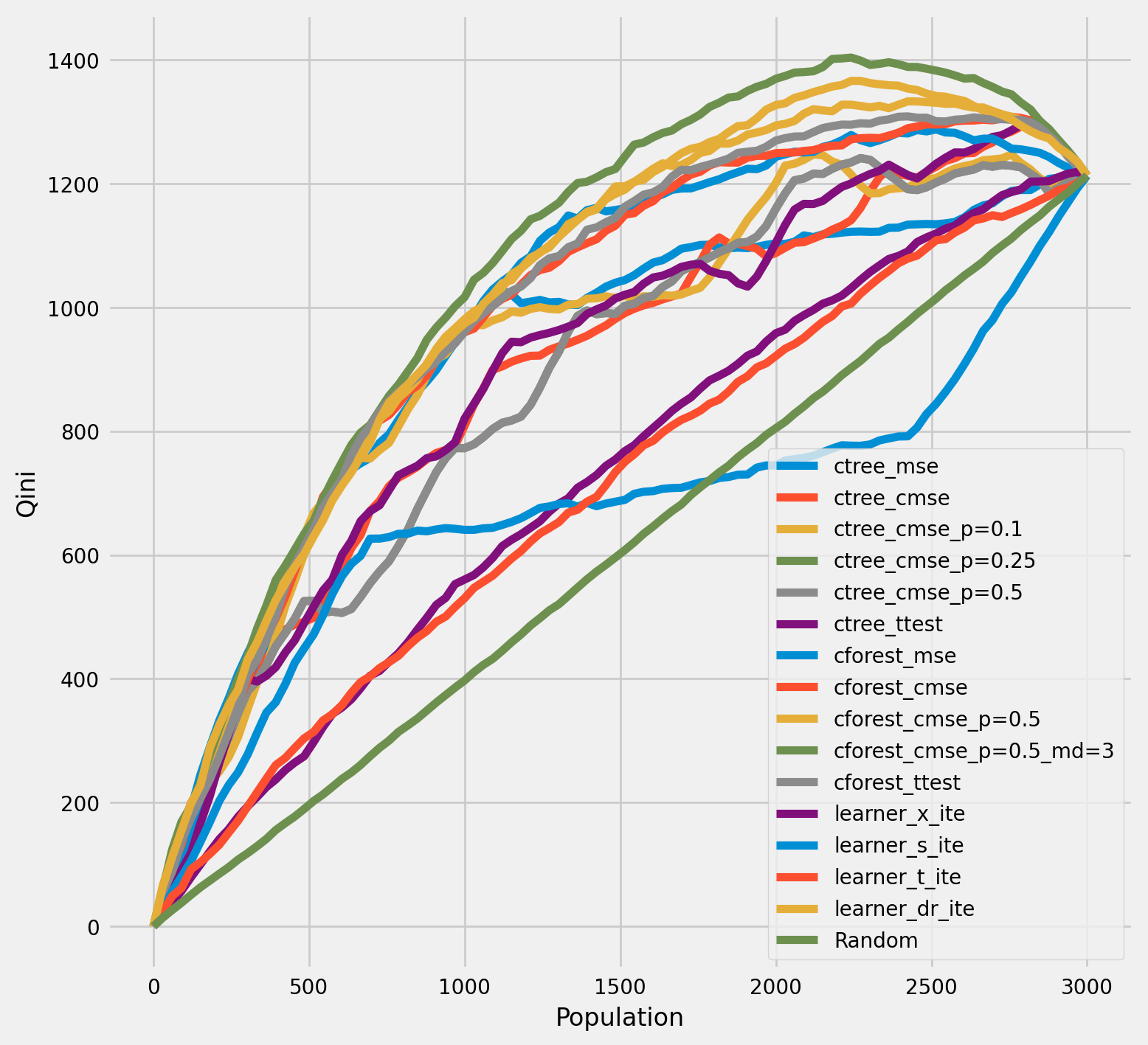
Qini chart¶
[33]:
plot_qini(df_result,
outcome_col='outcome',
treatment_col='is_treated',
treatment_effect_col='treatment_effect',
)
[34]:
df_qini = qini_score(df_result,
outcome_col='outcome',
treatment_col='is_treated',
treatment_effect_col='treatment_effect')
df_qini.sort_values(ascending=False)
[34]:
cforest_cmse_p=0.5_md=3 0.379598
learner_dr_ite 0.350053
cforest_cmse_p=0.5 0.344870
cforest_ttest 0.329592
cforest_mse 0.326770
cforest_cmse 0.323620
ctree_cmse_p=0.1 0.273112
ctree_mse 0.260141
ctree_ttest 0.247668
ctree_cmse 0.242333
ctree_cmse_p=0.25 0.231232
ctree_cmse_p=0.5 0.231232
learner_x_ite 0.096785
learner_t_ite 0.082948
learner_s_ite 0.055251
Random 0.000000
dtype: float64
Return outcomes along with estimated treatment effects¶
[35]:
ctree_outcomes = ctrees["ctree_mse"]["model"].predict(X_test, with_outcomes=True)
df_ctree_outcomes = pd.DataFrame(ctree_outcomes, columns=["Y0", "Y1", "ITE"])
df_ctree_outcomes.head()
[35]:
| Y0 | Y1 | ITE | |
|---|---|---|---|
| 0 | 0.434715 | 0.545400 | 0.110685 |
| 1 | 1.825205 | 2.008497 | 0.183293 |
| 2 | 0.453153 | 1.287316 | 0.834163 |
| 3 | 1.825205 | 2.008497 | 0.183293 |
| 4 | 1.825205 | 2.008497 | 0.183293 |
[36]:
cforest_outcomes = cforests["cforest_mse"]["model"].predict(X_test, with_outcomes=True)
df_cforest_outcomes = pd.DataFrame(cforest_outcomes, columns=["Y0", "Y1", "ITE"])
df_cforest_outcomes.head()
[36]:
| Y0 | Y1 | ITE | |
|---|---|---|---|
| 0 | 0.390998 | 0.575902 | 0.184904 |
| 1 | 1.445800 | 1.852422 | 0.406622 |
| 2 | 0.385520 | 0.982415 | 0.596895 |
| 3 | 1.279227 | 1.731512 | 0.452285 |
| 4 | 1.279100 | 1.730838 | 0.451738 |
Bootstrap confidence intervals for individual treatment effects¶
[37]:
alpha=0.05
tree = CausalTreeRegressor(criterion='causal_mse', control_name=0, min_samples_leaf=200, alpha=alpha)
[38]:
# For time measurements
for n_jobs in (4, mp.cpu_count() - 1):
for n_bootstraps in (10, 50, 100):
print(f"n_jobs: {n_jobs} n_bootstraps: {n_bootstraps}" )
tree.bootstrap_pool(
X=X,
treatment=w,
y=y,
n_bootstraps=n_bootstraps,
bootstrap_size=10000,
n_jobs=n_jobs,
verbose=False
)
n_jobs: 4 n_bootstraps: 10
100%|██████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████| 10/10 [00:00<00:00, 20.81it/s]
Function: bootstrap_pool Kwargs:{'n_bootstraps': 10, 'bootstrap_size': 10000, 'n_jobs': 4, 'verbose': False} Elapsed time: 0.9135
n_jobs: 4 n_bootstraps: 50
100%|██████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████| 50/50 [00:02<00:00, 21.29it/s]
Function: bootstrap_pool Kwargs:{'n_bootstraps': 50, 'bootstrap_size': 10000, 'n_jobs': 4, 'verbose': False} Elapsed time: 2.4252
n_jobs: 4 n_bootstraps: 100
100%|████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████| 100/100 [00:04<00:00, 21.70it/s]
Function: bootstrap_pool Kwargs:{'n_bootstraps': 100, 'bootstrap_size': 10000, 'n_jobs': 4, 'verbose': False} Elapsed time: 4.6932
n_jobs: 11 n_bootstraps: 10
100%|██████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████| 10/10 [00:00<00:00, 32.93it/s]
Function: bootstrap_pool Kwargs:{'n_bootstraps': 10, 'bootstrap_size': 10000, 'n_jobs': 11, 'verbose': False} Elapsed time: 0.6439
n_jobs: 11 n_bootstraps: 50
100%|██████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████| 50/50 [00:01<00:00, 30.42it/s]
Function: bootstrap_pool Kwargs:{'n_bootstraps': 50, 'bootstrap_size': 10000, 'n_jobs': 11, 'verbose': False} Elapsed time: 1.8508
n_jobs: 11 n_bootstraps: 100
100%|████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████| 100/100 [00:02<00:00, 33.67it/s]
Function: bootstrap_pool Kwargs:{'n_bootstraps': 100, 'bootstrap_size': 10000, 'n_jobs': 11, 'verbose': False} Elapsed time: 3.1553
[39]:
te, te_lower, te_upper = tree.fit_predict(
X=df_train[feature_names].values,
treatment=df_train["treatment"].values,
y=df_train["outcome"].values,
return_ci=True,
n_bootstraps=500,
bootstrap_size=5000,
n_jobs=mp.cpu_count() - 1,
verbose=False)
100%|████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████| 500/500 [00:06<00:00, 81.19it/s]
Function: bootstrap_pool Kwargs:{'n_bootstraps': 500, 'bootstrap_size': 5000, 'n_jobs': 11, 'verbose': False} Elapsed time: 6.4025
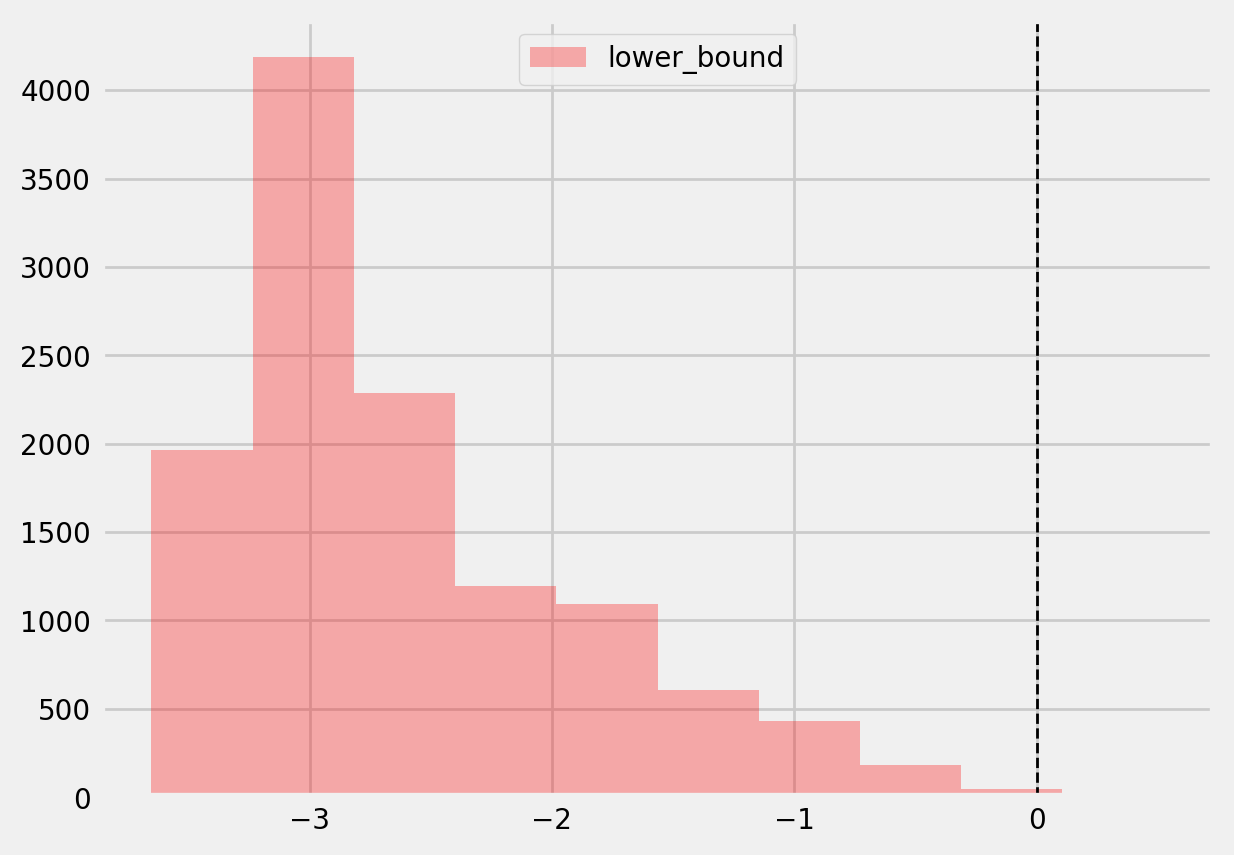
[40]:
plt.hist(te_lower, color='red', alpha=0.3, label='lower_bound')
plt.axvline(x = 0, color = 'black', linestyle='--', lw=1, label='')
plt.legend()
plt.show()
[41]:
# Significant estimates for negative and positive individual effects
# Default alpha = 0.05
bootstrap_neg = te[(te_lower < 0) & (te_upper < 0)]
bootstrap_pos = te[(te_lower > 0) & (te_upper > 0)]
print(bootstrap_neg.shape, bootstrap_pos.shape)
(0,) (7,)
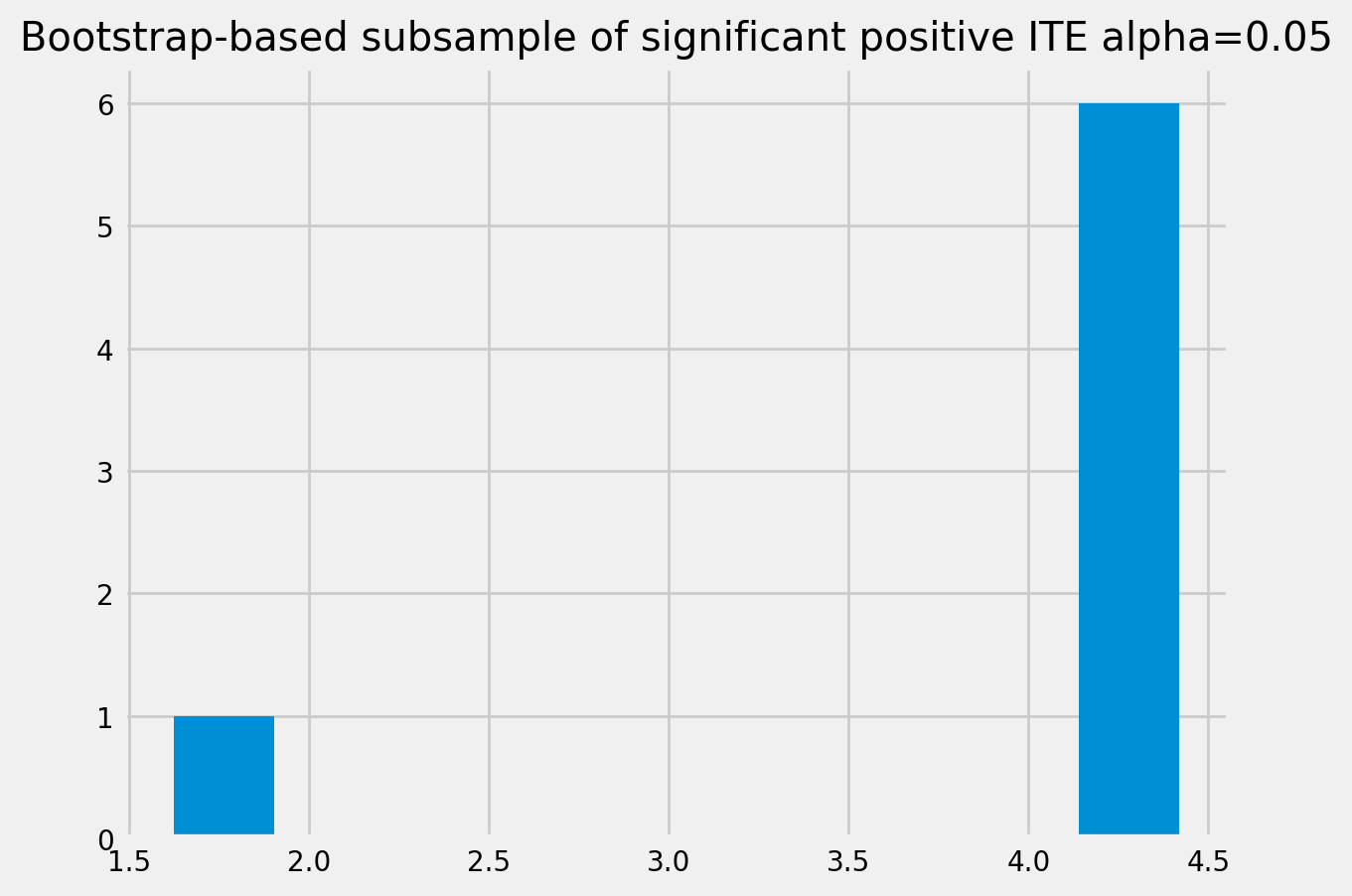
[42]:
plt.hist(bootstrap_neg)
plt.title(f'Bootstrap-based subsample of significant negative ITE. alpha={alpha}')
plt.show()
plt.hist(bootstrap_pos)
plt.title(f'Bootstrap-based subsample of significant positive ITE alpha={alpha}')
plt.show()
Average treatment effect¶
[43]:
tree = CausalTreeRegressor(criterion='causal_mse', control_name=0, min_samples_leaf=200, alpha=alpha)
te, te_lb, te_ub = tree.estimate_ate(X=X, treatment=w, y=y)
print('ATE:', te, 'Bounds:', (te_lb, te_ub ), 'alpha:', alpha)
ATE: 0.80899590297471 Bounds: (0.808752005241054, 0.8092398007083661) alpha: 0.05
CausalRandomForestRegressor ITE std¶
[44]:
crforest = CausalRandomForestRegressor(criterion="causal_mse", min_samples_leaf=200,
control_name=0, n_estimators=50, n_jobs=mp.cpu_count()-1)
crforest.fit(X=df_train[feature_names].values,
treatment=df_train['treatment'].values,
y=df_train['outcome'].values
)
[44]:
CausalRandomForestRegressor(min_samples_leaf=200, n_estimators=50, n_jobs=11)
[45]:
crforest_te_pred = crforest.predict(df_test[feature_names])
crforest_test_var = crforest.calculate_error(X_train=df_train[feature_names].values,
X_test=df_test[feature_names].values)
crforest_test_std = np.sqrt(crforest_test_var)
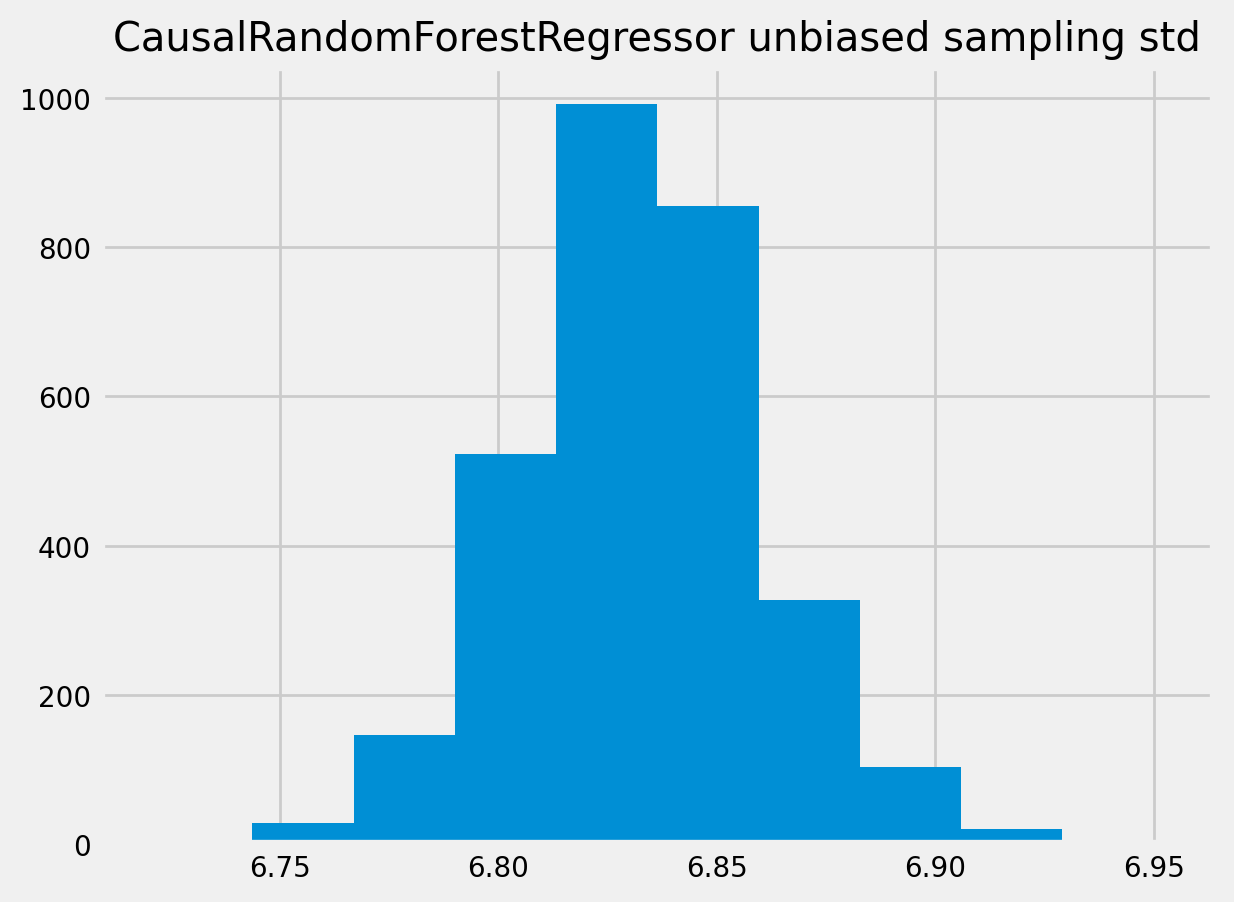
[46]:
plt.hist(crforest_test_std)
plt.title("CausalRandomForestRegressor unbiased sampling std")
plt.show()
[ ]:
[ ]: