Feature Selection for Uplift Trees by Zhao et al. (2020)
This notebook includes two sections:
Feature selection: demonstrate how to use Filter methods to select the most important numeric features
Performance evaluation: evaluate the AUUC performance with top features dataset
(Paper reference: Zhao, Zhenyu, et al. “Feature Selection Methods for Uplift Modeling.” arXiv preprint arXiv:2005.03447 (2020).)
[1]:
import numpy as np
import pandas as pd
[2]:
from causalml.dataset import make_uplift_classification
Import FilterSelect class for Filter methods
[3]:
from causalml.feature_selection.filters import FilterSelect
[4]:
from causalml.inference.tree import UpliftRandomForestClassifier
from causalml.inference.meta import BaseXRegressor, BaseRRegressor, BaseSRegressor, BaseTRegressor
from causalml.metrics import plot_gain, auuc_score
[5]:
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestRegressor
[6]:
import logging
logger = logging.getLogger('causalml')
logging.basicConfig(level=logging.INFO)
Generate dataset
Generate synthetic data using the built-in function.
[7]:
# define parameters for simulation
y_name = 'conversion'
treatment_group_keys = ['control', 'treatment1']
n = 10000
n_classification_features = 50
n_classification_informative = 10
n_classification_repeated = 0
n_uplift_increase_dict = {'treatment1': 8}
n_uplift_decrease_dict = {'treatment1': 4}
delta_uplift_increase_dict = {'treatment1': 0.1}
delta_uplift_decrease_dict = {'treatment1': -0.1}
random_seed = 20200808
[8]:
df, X_names = make_uplift_classification(
treatment_name=treatment_group_keys,
y_name=y_name,
n_samples=n,
n_classification_features=n_classification_features,
n_classification_informative=n_classification_informative,
n_classification_repeated=n_classification_repeated,
n_uplift_increase_dict=n_uplift_increase_dict,
n_uplift_decrease_dict=n_uplift_decrease_dict,
delta_uplift_increase_dict = delta_uplift_increase_dict,
delta_uplift_decrease_dict = delta_uplift_decrease_dict,
random_seed=random_seed
)
INFO:numexpr.utils:Note: NumExpr detected 12 cores but "NUMEXPR_MAX_THREADS" not set, so enforcing safe limit of 8.
INFO:numexpr.utils:NumExpr defaulting to 8 threads.
[9]:
df.head()
[9]:
| treatment_group_key | x1_informative | x2_informative | x3_informative | x4_informative | x5_informative | x6_informative | x7_informative | x8_informative | x9_informative | ... | x56_uplift_increase | x57_uplift_increase | x58_uplift_increase | x59_increase_mix | x60_uplift_decrease | x61_uplift_decrease | x62_uplift_decrease | x63_uplift_decrease | conversion | treatment_effect | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | treatment1 | -4.004496 | -1.250351 | -2.800557 | -0.368288 | -0.115549 | -2.492826 | 0.369516 | 0.290526 | 0.465153 | ... | 0.496144 | 1.847680 | -0.337894 | -0.672058 | 1.180352 | 0.778013 | 0.931000 | 2.947160 | 0 | 0 |
| 1 | treatment1 | -3.170028 | -0.135293 | 1.484246 | -2.131584 | -0.760103 | 1.764765 | 0.972124 | 1.407131 | -1.027603 | ... | 0.574955 | 3.578138 | 0.678118 | -0.545227 | -0.143942 | -0.015188 | 1.189643 | 1.943692 | 1 | 0 |
| 2 | treatment1 | -0.763386 | -0.785612 | 1.218781 | -0.725835 | 1.044489 | -1.521071 | -2.266684 | -1.614818 | -0.113647 | ... | 0.985076 | 1.079181 | 0.578092 | 0.574370 | -0.477429 | 0.679070 | 1.650897 | 2.768897 | 1 | 0 |
| 3 | control | 0.887727 | 0.049095 | -2.242776 | 1.530966 | 0.392623 | -0.203071 | -0.549329 | 0.107296 | -0.542277 | ... | -0.175352 | 0.683330 | 0.567545 | 0.349622 | -0.789203 | 2.315184 | 0.658607 | 1.734836 | 0 | 0 |
| 4 | control | -1.672922 | -1.156145 | 3.871476 | -1.883713 | -0.220122 | -4.615669 | 0.141980 | -0.933756 | -0.765592 | ... | 0.485798 | -0.355315 | 0.982488 | -0.007260 | 2.895155 | 0.261848 | -1.337001 | -0.639983 | 1 | 0 |
5 rows × 66 columns
[10]:
# Look at the conversion rate and sample size in each group
df.pivot_table(values='conversion',
index='treatment_group_key',
aggfunc=[np.mean, np.size],
margins=True)
[10]:
| mean | size | |
|---|---|---|
| conversion | conversion | |
| treatment_group_key | ||
| control | 0.50180 | 10000 |
| treatment1 | 0.59750 | 10000 |
| All | 0.54965 | 20000 |
[11]:
X_names
[11]:
['x1_informative',
'x2_informative',
'x3_informative',
'x4_informative',
'x5_informative',
'x6_informative',
'x7_informative',
'x8_informative',
'x9_informative',
'x10_informative',
'x11_irrelevant',
'x12_irrelevant',
'x13_irrelevant',
'x14_irrelevant',
'x15_irrelevant',
'x16_irrelevant',
'x17_irrelevant',
'x18_irrelevant',
'x19_irrelevant',
'x20_irrelevant',
'x21_irrelevant',
'x22_irrelevant',
'x23_irrelevant',
'x24_irrelevant',
'x25_irrelevant',
'x26_irrelevant',
'x27_irrelevant',
'x28_irrelevant',
'x29_irrelevant',
'x30_irrelevant',
'x31_irrelevant',
'x32_irrelevant',
'x33_irrelevant',
'x34_irrelevant',
'x35_irrelevant',
'x36_irrelevant',
'x37_irrelevant',
'x38_irrelevant',
'x39_irrelevant',
'x40_irrelevant',
'x41_irrelevant',
'x42_irrelevant',
'x43_irrelevant',
'x44_irrelevant',
'x45_irrelevant',
'x46_irrelevant',
'x47_irrelevant',
'x48_irrelevant',
'x49_irrelevant',
'x50_irrelevant',
'x51_uplift_increase',
'x52_uplift_increase',
'x53_uplift_increase',
'x54_uplift_increase',
'x55_uplift_increase',
'x56_uplift_increase',
'x57_uplift_increase',
'x58_uplift_increase',
'x59_increase_mix',
'x60_uplift_decrease',
'x61_uplift_decrease',
'x62_uplift_decrease',
'x63_uplift_decrease']
Feature selection with Filter methods
method = F (F Filter)
[12]:
filter_method = FilterSelect()
[13]:
# F Filter with order 1
method = 'F'
f_imp = filter_method.get_importance(df, X_names, y_name, method,
treatment_group = 'treatment1')
f_imp.head()
[13]:
| method | feature | rank | score | p_value | misc | |
|---|---|---|---|---|---|---|
| 0 | F filter | x53_uplift_increase | 1.0 | 190.321410 | 4.262512e-43 | df_num: 1.0, df_denom: 19996.0, order:1 |
| 0 | F filter | x57_uplift_increase | 2.0 | 127.136380 | 2.127676e-29 | df_num: 1.0, df_denom: 19996.0, order:1 |
| 0 | F filter | x3_informative | 3.0 | 66.273458 | 4.152970e-16 | df_num: 1.0, df_denom: 19996.0, order:1 |
| 0 | F filter | x4_informative | 4.0 | 59.407590 | 1.341417e-14 | df_num: 1.0, df_denom: 19996.0, order:1 |
| 0 | F filter | x62_uplift_decrease | 5.0 | 3.957507 | 4.667636e-02 | df_num: 1.0, df_denom: 19996.0, order:1 |
[14]:
# F Filter with order 2
method = 'F'
f_imp = filter_method.get_importance(df, X_names, y_name, method,
treatment_group = 'treatment1', order=2)
f_imp.head()
[14]:
| method | feature | rank | score | p_value | misc | |
|---|---|---|---|---|---|---|
| 0 | F filter | x53_uplift_increase | 1.0 | 107.368286 | 4.160720e-47 | df_num: 2.0, df_denom: 19994.0, order:2 |
| 0 | F filter | x57_uplift_increase | 2.0 | 70.138050 | 4.423736e-31 | df_num: 2.0, df_denom: 19994.0, order:2 |
| 0 | F filter | x3_informative | 3.0 | 36.499465 | 1.504356e-16 | df_num: 2.0, df_denom: 19994.0, order:2 |
| 0 | F filter | x4_informative | 4.0 | 31.780547 | 1.658731e-14 | df_num: 2.0, df_denom: 19994.0, order:2 |
| 0 | F filter | x55_uplift_increase | 5.0 | 27.494904 | 1.189886e-12 | df_num: 2.0, df_denom: 19994.0, order:2 |
[15]:
# F Filter with order 3
method = 'F'
f_imp = filter_method.get_importance(df, X_names, y_name, method,
treatment_group = 'treatment1', order=3)
f_imp.head()
[15]:
| method | feature | rank | score | p_value | misc | |
|---|---|---|---|---|---|---|
| 0 | F filter | x53_uplift_increase | 1.0 | 72.064224 | 2.373628e-46 | df_num: 3.0, df_denom: 19992.0, order:3 |
| 0 | F filter | x57_uplift_increase | 2.0 | 46.841718 | 3.710784e-30 | df_num: 3.0, df_denom: 19992.0, order:3 |
| 0 | F filter | x3_informative | 3.0 | 24.089980 | 1.484634e-15 | df_num: 3.0, df_denom: 19992.0, order:3 |
| 0 | F filter | x4_informative | 4.0 | 23.097310 | 6.414267e-15 | df_num: 3.0, df_denom: 19992.0, order:3 |
| 0 | F filter | x55_uplift_increase | 5.0 | 18.072880 | 1.044117e-11 | df_num: 3.0, df_denom: 19992.0, order:3 |
method = LR (likelihood ratio test)
[16]:
# LR Filter with order 1
method = 'LR'
lr_imp = filter_method.get_importance(df, X_names, y_name, method,
treatment_group = 'treatment1')
lr_imp.head()
[16]:
| method | feature | rank | score | p_value | misc | |
|---|---|---|---|---|---|---|
| 0 | LR filter | x53_uplift_increase | 1.0 | 203.811674 | 0.000000e+00 | df: 1, order: 1 |
| 0 | LR filter | x57_uplift_increase | 2.0 | 133.175328 | 0.000000e+00 | df: 1, order: 1 |
| 0 | LR filter | x3_informative | 3.0 | 64.366711 | 9.992007e-16 | df: 1, order: 1 |
| 0 | LR filter | x4_informative | 4.0 | 52.389798 | 4.550804e-13 | df: 1, order: 1 |
| 0 | LR filter | x62_uplift_decrease | 5.0 | 4.064347 | 4.379760e-02 | df: 1, order: 1 |
[17]:
# LR Filter with order 2
method = 'LR'
lr_imp = filter_method.get_importance(df, X_names, y_name, method,
treatment_group = 'treatment1',order=2)
lr_imp.head()
[17]:
| method | feature | rank | score | p_value | misc | |
|---|---|---|---|---|---|---|
| 0 | LR filter | x53_uplift_increase | 1.0 | 277.639095 | 0.000000e+00 | df: 2, order: 2 |
| 0 | LR filter | x57_uplift_increase | 2.0 | 156.134112 | 0.000000e+00 | df: 2, order: 2 |
| 0 | LR filter | x55_uplift_increase | 3.0 | 71.478979 | 3.330669e-16 | df: 2, order: 2 |
| 0 | LR filter | x3_informative | 4.0 | 44.938973 | 1.744319e-10 | df: 2, order: 2 |
| 0 | LR filter | x4_informative | 5.0 | 29.179971 | 4.609458e-07 | df: 2, order: 2 |
[18]:
# LR Filter with order 3
method = 'LR'
lr_imp = filter_method.get_importance(df, X_names, y_name, method,
treatment_group = 'treatment1',order=3)
lr_imp.head()
[18]:
| method | feature | rank | score | p_value | misc | |
|---|---|---|---|---|---|---|
| 0 | LR filter | x53_uplift_increase | 1.0 | 290.389201 | 0.000000e+00 | df: 3, order: 3 |
| 0 | LR filter | x57_uplift_increase | 2.0 | 153.942614 | 0.000000e+00 | df: 3, order: 3 |
| 0 | LR filter | x55_uplift_increase | 3.0 | 70.626667 | 3.108624e-15 | df: 3, order: 3 |
| 0 | LR filter | x3_informative | 4.0 | 45.477851 | 7.323235e-10 | df: 3, order: 3 |
| 0 | LR filter | x4_informative | 5.0 | 30.466528 | 1.100881e-06 | df: 3, order: 3 |
method = KL (KL divergence)
[19]:
method = 'KL'
kl_imp = filter_method.get_importance(df, X_names, y_name, method,
treatment_group = 'treatment1',
n_bins=10)
kl_imp.head()
[19]:
| method | feature | rank | score | p_value | misc | |
|---|---|---|---|---|---|---|
| 0 | KL filter | x53_uplift_increase | 1.0 | 0.022997 | None | number_of_bins: 10 |
| 0 | KL filter | x57_uplift_increase | 2.0 | 0.014884 | None | number_of_bins: 10 |
| 0 | KL filter | x4_informative | 3.0 | 0.012103 | None | number_of_bins: 10 |
| 0 | KL filter | x3_informative | 4.0 | 0.010179 | None | number_of_bins: 10 |
| 0 | KL filter | x55_uplift_increase | 5.0 | 0.003836 | None | number_of_bins: 10 |
We found all these 3 filter methods were able to rank most of the informative and uplift increase features on the top.
Performance evaluation
Evaluate the AUUC (Area Under the Uplift Curve) score with several uplift models when using top features dataset
[20]:
# train test split
df_train, df_test = train_test_split(df, test_size=0.2, random_state=111)
[21]:
# convert treatment column to 1 (treatment1) and 0 (control)
treatments = np.where((df_test['treatment_group_key']=='treatment1'), 1, 0)
print(treatments[:10])
print(df_test['treatment_group_key'][:10])
[0 0 1 1 0 1 1 0 0 0]
18998 control
11536 control
8552 treatment1
2652 treatment1
19671 control
13244 treatment1
3075 treatment1
8746 control
18530 control
5066 control
Name: treatment_group_key, dtype: object
Uplift RandomForest Classfier
[22]:
uplift_model = UpliftRandomForestClassifier(control_name='control', max_depth=8)
[23]:
# using all features
features = X_names
uplift_model.fit(X = df_train[features].values,
treatment = df_train['treatment_group_key'].values,
y = df_train[y_name].values)
y_preds = uplift_model.predict(df_test[features].values)
Select top N features based on KL filter
[24]:
top_n = 10
top_10_features = kl_imp['feature'][:top_n]
print(top_10_features)
0 x53_uplift_increase
0 x57_uplift_increase
0 x4_informative
0 x3_informative
0 x55_uplift_increase
0 x1_informative
0 x56_uplift_increase
0 x51_uplift_increase
0 x38_irrelevant
0 x58_uplift_increase
Name: feature, dtype: object
[25]:
top_n = 15
top_15_features = kl_imp['feature'][:top_n]
print(top_15_features)
0 x53_uplift_increase
0 x57_uplift_increase
0 x4_informative
0 x3_informative
0 x55_uplift_increase
0 x1_informative
0 x56_uplift_increase
0 x51_uplift_increase
0 x38_irrelevant
0 x58_uplift_increase
0 x48_irrelevant
0 x15_irrelevant
0 x27_irrelevant
0 x62_uplift_decrease
0 x23_irrelevant
Name: feature, dtype: object
[26]:
top_n = 20
top_20_features = kl_imp['feature'][:top_n]
print(top_20_features)
0 x53_uplift_increase
0 x57_uplift_increase
0 x4_informative
0 x3_informative
0 x55_uplift_increase
0 x1_informative
0 x56_uplift_increase
0 x51_uplift_increase
0 x38_irrelevant
0 x58_uplift_increase
0 x48_irrelevant
0 x15_irrelevant
0 x27_irrelevant
0 x62_uplift_decrease
0 x23_irrelevant
0 x29_irrelevant
0 x6_informative
0 x45_irrelevant
0 x40_irrelevant
0 x25_irrelevant
Name: feature, dtype: object
Train the Uplift model again with top N features
[27]:
# using top 10 features
features = top_10_features
uplift_model.fit(X = df_train[features].values,
treatment = df_train['treatment_group_key'].values,
y = df_train[y_name].values)
y_preds_t10 = uplift_model.predict(df_test[features].values)
[28]:
# using top 15 features
features = top_15_features
uplift_model.fit(X = df_train[features].values,
treatment = df_train['treatment_group_key'].values,
y = df_train[y_name].values)
y_preds_t15 = uplift_model.predict(df_test[features].values)
[29]:
# using top 20 features
features = top_20_features
uplift_model.fit(X = df_train[features].values,
treatment = df_train['treatment_group_key'].values,
y = df_train[y_name].values)
y_preds_t20 = uplift_model.predict(df_test[features].values)
Print results for Uplift model
[30]:
df_preds = pd.DataFrame([y_preds.ravel(),
y_preds_t10.ravel(),
y_preds_t15.ravel(),
y_preds_t20.ravel(),
treatments,
df_test[y_name].ravel()],
index=['All', 'Top 10', 'Top 15', 'Top 20', 'is_treated', y_name]).T
plot_gain(df_preds, outcome_col=y_name, treatment_col='is_treated')
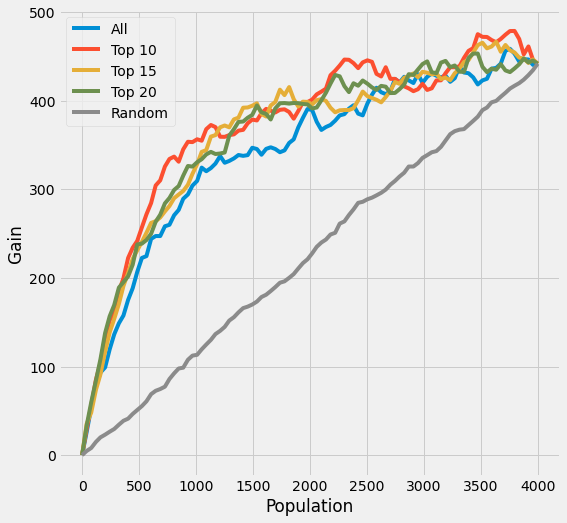
[31]:
auuc_score(df_preds, outcome_col=y_name, treatment_col='is_treated')
[31]:
All 0.773405
Top 10 0.841204
Top 15 0.816100
Top 20 0.816252
Random 0.506801
dtype: float64
R Learner as base and feed in Random Forest Regressor
[32]:
r_rf_learner = BaseRRegressor(
RandomForestRegressor(
n_estimators = 100,
max_depth = 8,
min_samples_leaf = 100
),
control_name='control')
[33]:
# using all features
features = X_names
r_rf_learner.fit(X = df_train[features].values,
treatment = df_train['treatment_group_key'].values,
y = df_train[y_name].values)
y_preds = r_rf_learner.predict(df_test[features].values)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment1 with R-loss
[34]:
# using top 10 features
features = top_10_features
r_rf_learner.fit(X = df_train[features].values,
treatment = df_train['treatment_group_key'].values,
y = df_train[y_name].values)
y_preds_t10 = r_rf_learner.predict(df_test[features].values)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment1 with R-loss
[35]:
# using top 15 features
features = top_15_features
r_rf_learner.fit(X = df_train[features].values,
treatment = df_train['treatment_group_key'].values,
y = df_train[y_name].values)
y_preds_t15 = r_rf_learner.predict(df_test[features].values)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment1 with R-loss
[36]:
# using top 20 features
features = top_20_features
r_rf_learner.fit(X = df_train[features].values,
treatment = df_train['treatment_group_key'].values,
y = df_train[y_name].values)
y_preds_t20 = r_rf_learner.predict(df_test[features].values)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment1 with R-loss
Print results for R Learner
[37]:
df_preds = pd.DataFrame([y_preds.ravel(),
y_preds_t10.ravel(),
y_preds_t15.ravel(),
y_preds_t20.ravel(),
treatments,
df_test[y_name].ravel()],
index=['All', 'Top 10', 'Top 15', 'Top 20', 'is_treated', y_name]).T
plot_gain(df_preds, outcome_col=y_name, treatment_col='is_treated')
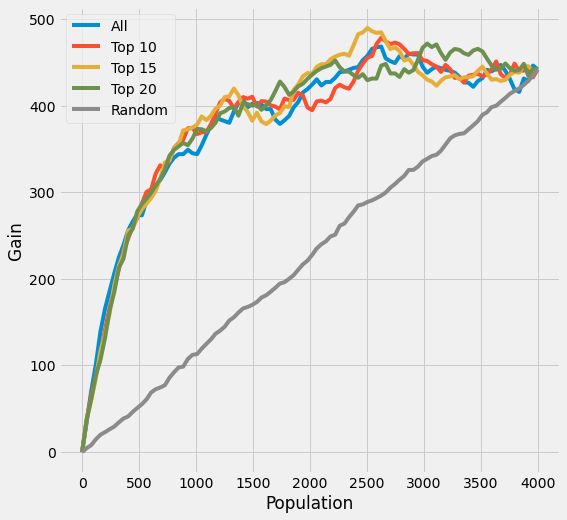
[38]:
# print out AUUC score
auuc_score(df_preds, outcome_col=y_name, treatment_col='is_treated')
[38]:
All 0.859891
Top 10 0.865159
Top 15 0.872650
Top 20 0.870669
Random 0.506801
dtype: float64
(a relatively smaller enhancement on the AUUC is observed in this R Learner case)
S Learner as base and feed in Random Forest Regressor
[39]:
slearner_rf = BaseSRegressor(
RandomForestRegressor(
n_estimators = 100,
max_depth = 8,
min_samples_leaf = 100
),
control_name='control')
[40]:
# using all features
features = X_names
slearner_rf.fit(X = df_train[features].values,
treatment = df_train['treatment_group_key'].values,
y = df_train[y_name].values)
y_preds = slearner_rf.predict(df_test[features].values)
[41]:
# using top 10 features
features = top_10_features
slearner_rf.fit(X = df_train[features].values,
treatment = df_train['treatment_group_key'].values,
y = df_train[y_name].values)
y_preds_t10 = slearner_rf.predict(df_test[features].values)
[42]:
# using top 15 features
features = top_15_features
slearner_rf.fit(X = df_train[features].values,
treatment = df_train['treatment_group_key'].values,
y = df_train[y_name].values)
y_preds_t15 = slearner_rf.predict(df_test[features].values)
[43]:
# using top 20 features
features = top_20_features
slearner_rf.fit(X = df_train[features].values,
treatment = df_train['treatment_group_key'].values,
y = df_train[y_name].values)
y_preds_t20 = slearner_rf.predict(df_test[features].values)
Print results for S Learner
[44]:
df_preds = pd.DataFrame([y_preds.ravel(),
y_preds_t10.ravel(),
y_preds_t15.ravel(),
y_preds_t20.ravel(),
treatments,
df_test[y_name].ravel()],
index=['All', 'Top 10', 'Top 15', 'Top 20', 'is_treated', y_name]).T
plot_gain(df_preds, outcome_col=y_name, treatment_col='is_treated')
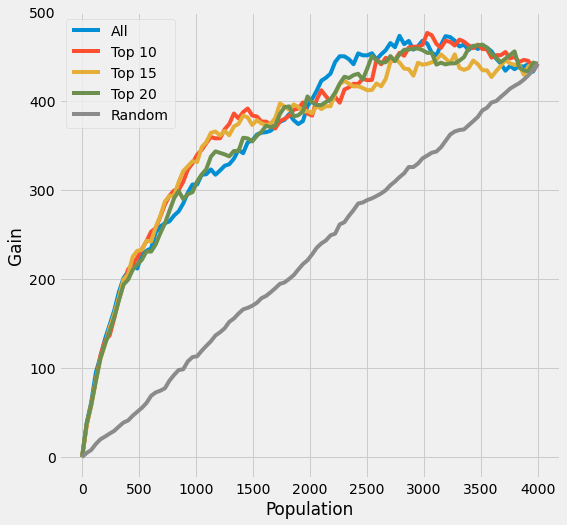
[45]:
# print out AUUC score
auuc_score(df_preds, outcome_col=y_name, treatment_col='is_treated')
[45]:
All 0.824483
Top 10 0.832872
Top 15 0.817835
Top 20 0.816149
Random 0.506801
dtype: float64
In this notebook, we demonstrated how our Filter method functions are able to select important features and enhance the AUUC performance (while the results might vary among different datasets, models and hyper-parameters).
[ ]: