Meta-Learners Examples - Single/Multiple Treatment Cases
This notebook only contains regression examples.
[1]:
%reload_ext autoreload
%autoreload 2
[2]:
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
from sklearn.linear_model import LinearRegression, LogisticRegression
from sklearn.model_selection import train_test_split
import statsmodels.api as sm
from xgboost import XGBRegressor, XGBClassifier
import warnings
# from causalml.inference.meta import XGBTLearner, MLPTLearner
from causalml.inference.meta import BaseSRegressor, BaseTRegressor, BaseXRegressor, BaseRRegressor
from causalml.inference.meta import BaseSClassifier, BaseTClassifier, BaseXClassifier, BaseRClassifier
from causalml.inference.meta import LRSRegressor
from causalml.match import NearestNeighborMatch, MatchOptimizer, create_table_one
from causalml.propensity import ElasticNetPropensityModel
from causalml.dataset import *
from causalml.metrics import *
warnings.filterwarnings('ignore')
plt.style.use('fivethirtyeight')
pd.set_option('display.float_format', lambda x: '%.4f' % x)
# imports from package
import logging
from sklearn.dummy import DummyRegressor
from sklearn.metrics import mean_squared_error as mse
from sklearn.metrics import mean_absolute_error as mae
import statsmodels.api as sm
from copy import deepcopy
logger = logging.getLogger('causalml')
logging.basicConfig(level=logging.INFO)
%matplotlib inline
/Users/jeong/.conda/envs/py36/lib/python3.6/site-packages/sklearn/utils/deprecation.py:144: FutureWarning: The sklearn.utils.testing module is deprecated in version 0.22 and will be removed in version 0.24. The corresponding classes / functions should instead be imported from sklearn.utils. Anything that cannot be imported from sklearn.utils is now part of the private API.
warnings.warn(message, FutureWarning)
Single Treatment Case
Generate synthetic data
[3]:
# Generate synthetic data using mode 1
y, X, treatment, tau, b, e = synthetic_data(mode=1, n=10000, p=8, sigma=1.0)
treatment = np.array(['treatment_a' if val==1 else 'control' for val in treatment])
S-Learner
ATE
[4]:
learner_s = BaseSRegressor(XGBRegressor(), control_name='control')
ate_s = learner_s.estimate_ate(X=X, treatment=treatment, y=y, return_ci=False, bootstrap_ci=False)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.6622
INFO:causalml: RMSE (Treatment): 0.6941
INFO:causalml: sMAPE (Control): 0.6536
INFO:causalml: sMAPE (Treatment): 0.3721
INFO:causalml: Gini (Control): 0.8248
INFO:causalml: Gini (Treatment): 0.8156
[5]:
ate_s
[5]:
array([0.57431368])
ATE w/ Confidence Intervals
[6]:
alpha = 0.05
learner_s = BaseSRegressor(XGBRegressor(), ate_alpha=alpha, control_name='control')
ate_s, ate_s_lb, ate_s_ub = learner_s.estimate_ate(X=X, treatment=treatment, y=y, return_ci=True,
bootstrap_ci=False)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.6622
INFO:causalml: RMSE (Treatment): 0.6941
INFO:causalml: sMAPE (Control): 0.6536
INFO:causalml: sMAPE (Treatment): 0.3721
INFO:causalml: Gini (Control): 0.8248
INFO:causalml: Gini (Treatment): 0.8156
[7]:
np.vstack((ate_s_lb, ate_s, ate_s_ub))
[7]:
array([[0.54689052],
[0.57431368],
[0.60173684]])
ATE w/ Boostrap Confidence Intervals
[8]:
ate_s_b, ate_s_lb_b, ate_s_ub_b = learner_s.estimate_ate(X=X, treatment=treatment, y=y, return_ci=True,
bootstrap_ci=True, n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.6622
INFO:causalml: RMSE (Treatment): 0.6941
INFO:causalml: sMAPE (Control): 0.6536
INFO:causalml: sMAPE (Treatment): 0.3721
INFO:causalml: Gini (Control): 0.8248
INFO:causalml: Gini (Treatment): 0.8156
INFO:causalml:Bootstrap Confidence Intervals for ATE
100%|██████████| 100/100 [01:14<00:00, 1.34it/s]
[9]:
np.vstack((ate_s_lb_b, ate_s_b, ate_s_ub_b))
[9]:
array([[0.51141982],
[0.57431368],
[0.64097547]])
CATE
[10]:
learner_s = BaseSRegressor(XGBRegressor(), control_name='control')
cate_s = learner_s.fit_predict(X=X, treatment=treatment, y=y, return_ci=False)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.6622
INFO:causalml: RMSE (Treatment): 0.6941
INFO:causalml: sMAPE (Control): 0.6536
INFO:causalml: sMAPE (Treatment): 0.3721
INFO:causalml: Gini (Control): 0.8248
INFO:causalml: Gini (Treatment): 0.8156
[11]:
cate_s
[11]:
array([[0.37674308],
[0.42519259],
[0.60864675],
...,
[0.19940662],
[0.35013032],
[0.78372002]])
CATE w/ Confidence Intervals
[12]:
alpha = 0.05
learner_s = BaseSRegressor(XGBRegressor(), ate_alpha=alpha, control_name='control')
cate_s, cate_s_lb, cate_s_ub = learner_s.fit_predict(X=X, treatment=treatment, y=y, return_ci=True,
n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.6622
INFO:causalml: RMSE (Treatment): 0.6941
INFO:causalml: sMAPE (Control): 0.6536
INFO:causalml: sMAPE (Treatment): 0.3721
INFO:causalml: Gini (Control): 0.8248
INFO:causalml: Gini (Treatment): 0.8156
INFO:causalml:Bootstrap Confidence Intervals
100%|██████████| 100/100 [01:02<00:00, 1.59it/s]
[13]:
cate_s
[13]:
array([[0.37674308],
[0.42519259],
[0.60864675],
...,
[0.19940662],
[0.35013032],
[0.78372002]])
[14]:
cate_s_lb
[14]:
array([[-0.18972662],
[ 0.20548496],
[ 0.09983036],
...,
[-0.62837307],
[-0.19766161],
[-0.07736247]])
[15]:
cate_s_ub
[15]:
array([[0.8139405 ],
[1.278447 ],
[1.21720439],
...,
[0.90244564],
[0.9450083 ],
[1.1529291 ]])
T-Learner
ATE w/ Confidence Intervals
[16]:
learner_t = BaseTRegressor(XGBRegressor(), control_name='control')
ate_t, ate_t_lb, ate_t_ub = learner_t.estimate_ate(X=X, treatment=treatment, y=y)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4868
INFO:causalml: RMSE (Treatment): 0.5434
INFO:causalml: sMAPE (Control): 0.5230
INFO:causalml: sMAPE (Treatment): 0.3114
INFO:causalml: Gini (Control): 0.9216
INFO:causalml: Gini (Treatment): 0.8988
[17]:
np.vstack((ate_t_lb, ate_t, ate_t_ub))
[17]:
array([[0.55534845],
[0.58090983],
[0.60647121]])
ATE w/ Boostrap Confidence Intervals
[18]:
ate_t_b, ate_t_lb_b, ate_t_ub_b = learner_t.estimate_ate(X=X, treatment=treatment, y=y, bootstrap_ci=True,
n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4868
INFO:causalml: RMSE (Treatment): 0.5434
INFO:causalml: sMAPE (Control): 0.5230
INFO:causalml: sMAPE (Treatment): 0.3114
INFO:causalml: Gini (Control): 0.9216
INFO:causalml: Gini (Treatment): 0.8988
INFO:causalml:Bootstrap Confidence Intervals for ATE
100%|██████████| 100/100 [01:00<00:00, 1.66it/s]
[19]:
np.vstack((ate_t_lb_b, ate_t_b, ate_t_ub_b))
[19]:
array([[0.51343277],
[0.58090983],
[0.65843097]])
CATE
[20]:
learner_t = BaseTRegressor(XGBRegressor(), control_name='control')
cate_t = learner_t.fit_predict(X=X, treatment=treatment, y=y)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4868
INFO:causalml: RMSE (Treatment): 0.5434
INFO:causalml: sMAPE (Control): 0.5230
INFO:causalml: sMAPE (Treatment): 0.3114
INFO:causalml: Gini (Control): 0.9216
INFO:causalml: Gini (Treatment): 0.8988
[21]:
cate_t
[21]:
array([[ 0.23669004],
[-0.0793891 ],
[-0.10774326],
...,
[ 0.30539629],
[ 0.50784194],
[ 0.00356007]])
CATE w/ Confidence Intervals
[22]:
learner_t = BaseTRegressor(XGBRegressor(), control_name='control')
cate_t, cate_t_lb, cate_t_ub = learner_t.fit_predict(X=X, treatment=treatment, y=y, return_ci=True, n_bootstraps=100,
bootstrap_size=5000)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4868
INFO:causalml: RMSE (Treatment): 0.5434
INFO:causalml: sMAPE (Control): 0.5230
INFO:causalml: sMAPE (Treatment): 0.3114
INFO:causalml: Gini (Control): 0.9216
INFO:causalml: Gini (Treatment): 0.8988
INFO:causalml:Bootstrap Confidence Intervals
100%|██████████| 100/100 [00:59<00:00, 1.68it/s]
[23]:
cate_t
[23]:
array([[ 0.23669004],
[-0.0793891 ],
[-0.10774326],
...,
[ 0.30539629],
[ 0.50784194],
[ 0.00356007]])
[24]:
cate_t_lb
[24]:
array([[-0.6752711 ],
[-0.72038152],
[-1.2330182 ],
...,
[-0.82131582],
[-0.48846376],
[-0.39046848]])
[25]:
cate_t_ub
[25]:
array([[1.66480025],
[1.60697527],
[2.06829221],
...,
[1.64941401],
[1.59083122],
[1.53139764]])
X-Learner
ATE w/ Confidence Intervals
With Propensity Score Input
[26]:
learner_x = BaseXRegressor(XGBRegressor(), control_name='control')
ate_x, ate_x_lb, ate_x_ub = learner_x.estimate_ate(X=X, treatment=treatment, y=y, p=e)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4868
INFO:causalml: RMSE (Treatment): 0.5434
INFO:causalml: sMAPE (Control): 0.5230
INFO:causalml: sMAPE (Treatment): 0.3114
INFO:causalml: Gini (Control): 0.9216
INFO:causalml: Gini (Treatment): 0.8988
[27]:
np.vstack((ate_x_lb, ate_x, ate_x_ub))
[27]:
array([[0.51454586],
[0.53721713],
[0.55988839]])
Without Propensity Score input
[28]:
ate_x_no_p, ate_x_lb_no_p, ate_x_ub_no_p = learner_x.estimate_ate(X=X, treatment=treatment, y=y)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4868
INFO:causalml: RMSE (Treatment): 0.5434
INFO:causalml: sMAPE (Control): 0.5230
INFO:causalml: sMAPE (Treatment): 0.3114
INFO:causalml: Gini (Control): 0.9216
INFO:causalml: Gini (Treatment): 0.8988
[29]:
np.vstack((ate_x_lb_no_p, ate_x_no_p, ate_x_ub_no_p))
[29]:
array([[0.51334384],
[0.53600211],
[0.55866038]])
[30]:
learner_x.propensity_model
[30]:
{'treatment_a': {'all training': LogisticRegressionCV(Cs=array([1.00230524, 2.15608891, 4.63802765, 9.97700064]),
class_weight=None,
cv=StratifiedKFold(n_splits=3, random_state=None, shuffle=True),
dual=False, fit_intercept=True, intercept_scaling=1.0,
l1_ratios=array([0.001 , 0.33366667, 0.66633333, 0.999 ]),
max_iter=100, multi_class='auto', n_jobs=None,
penalty='elasticnet', random_state=None, refit=True,
scoring=None, solver='saga', tol=0.0001, verbose=0)}}
ATE w/ Boostrap Confidence Intervals
With Propensity Score Input
[31]:
ate_x_b, ate_x_lb_b, ate_x_ub_b = learner_x.estimate_ate(X=X, treatment=treatment, y=y, p=e, bootstrap_ci=True,
n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4868
INFO:causalml: RMSE (Treatment): 0.5434
INFO:causalml: sMAPE (Control): 0.5230
INFO:causalml: sMAPE (Treatment): 0.3114
INFO:causalml: Gini (Control): 0.9216
INFO:causalml: Gini (Treatment): 0.8988
INFO:causalml:Bootstrap Confidence Intervals for ATE
100%|██████████| 100/100 [01:55<00:00, 1.15s/it]
[32]:
np.vstack((ate_x_lb_b, ate_x_b, ate_x_ub_b))
[32]:
array([[0.46262759],
[0.53721713],
[0.59662513]])
Without Propensity Score Input
[33]:
ate_x_b_no_p, ate_x_lb_b_no_p, ate_x_ub_b_no_p = learner_x.estimate_ate(X=X, treatment=treatment, y=y, bootstrap_ci=True,
n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4868
INFO:causalml: RMSE (Treatment): 0.5434
INFO:causalml: sMAPE (Control): 0.5230
INFO:causalml: sMAPE (Treatment): 0.3114
INFO:causalml: Gini (Control): 0.9216
INFO:causalml: Gini (Treatment): 0.8988
INFO:causalml:Bootstrap Confidence Intervals for ATE
100%|██████████| 100/100 [01:56<00:00, 1.17s/it]
[34]:
np.vstack((ate_x_lb_b_no_p, ate_x_b_no_p, ate_x_ub_b_no_p))
[34]:
array([[0.44360865],
[0.53598752],
[0.59794413]])
CATE
With Propensity Score Input
[35]:
learner_x = BaseXRegressor(XGBRegressor(), control_name='control')
cate_x = learner_x.fit_predict(X=X, treatment=treatment, y=y, p=e)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4868
INFO:causalml: RMSE (Treatment): 0.5434
INFO:causalml: sMAPE (Control): 0.5230
INFO:causalml: sMAPE (Treatment): 0.3114
INFO:causalml: Gini (Control): 0.9216
INFO:causalml: Gini (Treatment): 0.8988
[36]:
cate_x
[36]:
array([[0.05178452],
[0.01907274],
[0.79584839],
...,
[0.18147876],
[0.34742898],
[0.23145415]])
Without Propensity Score Input
[37]:
cate_x_no_p = learner_x.fit_predict(X=X, treatment=treatment, y=y)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4868
INFO:causalml: RMSE (Treatment): 0.5434
INFO:causalml: sMAPE (Control): 0.5230
INFO:causalml: sMAPE (Treatment): 0.3114
INFO:causalml: Gini (Control): 0.9216
INFO:causalml: Gini (Treatment): 0.8988
[38]:
cate_x_no_p
[38]:
array([[0.06426511],
[0.0189166 ],
[0.78233515],
...,
[0.2237187 ],
[0.29647103],
[0.2359861 ]])
CATE w/ Confidence Intervals
With Propensity Score Input
[39]:
learner_x = BaseXRegressor(XGBRegressor(), control_name='control')
cate_x, cate_x_lb, cate_x_ub = learner_x.fit_predict(X=X, treatment=treatment, y=y, p=e, return_ci=True,
n_bootstraps=100, bootstrap_size=3000)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4868
INFO:causalml: RMSE (Treatment): 0.5434
INFO:causalml: sMAPE (Control): 0.5230
INFO:causalml: sMAPE (Treatment): 0.3114
INFO:causalml: Gini (Control): 0.9216
INFO:causalml: Gini (Treatment): 0.8988
INFO:causalml:Bootstrap Confidence Intervals
100%|██████████| 100/100 [01:14<00:00, 1.34it/s]
[40]:
cate_x
[40]:
array([[0.05178452],
[0.01907274],
[0.79584839],
...,
[0.18147876],
[0.34742898],
[0.23145415]])
[41]:
cate_x_lb
[41]:
array([[-0.71763188],
[-0.79487709],
[-0.329782 ],
...,
[-0.57672694],
[-0.48450804],
[-0.43157597]])
[42]:
cate_x_ub
[42]:
array([[1.40320321],
[1.59906792],
[1.59324502],
...,
[1.07747513],
[1.30836353],
[1.18985624]])
Without Propensity Score Input
[43]:
cate_x_no_p, cate_x_lb_no_p, cate_x_ub_no_p = learner_x.fit_predict(X=X, treatment=treatment, y=y, return_ci=True,
n_bootstraps=100, bootstrap_size=3000)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4868
INFO:causalml: RMSE (Treatment): 0.5434
INFO:causalml: sMAPE (Control): 0.5230
INFO:causalml: sMAPE (Treatment): 0.3114
INFO:causalml: Gini (Control): 0.9216
INFO:causalml: Gini (Treatment): 0.8988
INFO:causalml:Bootstrap Confidence Intervals
100%|██████████| 100/100 [01:16<00:00, 1.31it/s]
[44]:
cate_x_no_p
[44]:
array([[0.06430496],
[0.01891659],
[0.78209735],
...,
[0.22376976],
[0.29645377],
[0.23597794]])
[45]:
cate_x_lb_no_p
[45]:
array([[-0.62013372],
[-0.90236405],
[-0.31043938],
...,
[-0.54219561],
[-0.2852425 ],
[-0.37437315]])
[46]:
cate_x_ub_no_p
[46]:
array([[1.4199368 ],
[1.45096372],
[1.57656827],
...,
[1.34583137],
[1.37899369],
[1.25074382]])
R-Learner
ATE w/ Confidence Intervals
With Propensity Score Input
[47]:
learner_r = BaseRRegressor(XGBRegressor(), control_name='control')
ate_r, ate_r_lb, ate_r_ub = learner_r.estimate_ate(X=X, treatment=treatment, y=y, p=e)
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
[48]:
np.vstack((ate_r_lb, ate_r, ate_r_ub))
[48]:
array([[0.55904178],
[0.55951123],
[0.55998069]])
Without Propensity Score Input
[49]:
ate_r_no_p, ate_r_lb_no_p, ate_r_ub_no_p = learner_r.estimate_ate(X=X, treatment=treatment, y=y)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
[50]:
np.vstack((ate_r_lb_no_p, ate_r_no_p, ate_r_ub_no_p))
[50]:
array([[0.49307912],
[0.49354918],
[0.49401924]])
[51]:
learner_r.propensity_model
[51]:
{'treatment_a': {'all training': LogisticRegressionCV(Cs=array([1.00230524, 2.15608891, 4.63802765, 9.97700064]),
class_weight=None,
cv=KFold(n_splits=5, random_state=None, shuffle=True),
dual=False, fit_intercept=True, intercept_scaling=1.0,
l1_ratios=array([0.001 , 0.33366667, 0.66633333, 0.999 ]),
max_iter=100, multi_class='auto', n_jobs=None,
penalty='elasticnet', random_state=None, refit=True,
scoring=None, solver='saga', tol=0.0001, verbose=0)}}
ATE w/ Boostrap Confidence Intervals
With Propensity Score Input
[52]:
ate_r_b, ate_r_lb_b, ate_r_ub_b = learner_r.estimate_ate(X=X, treatment=treatment, y=y, p=e, bootstrap_ci=True,
n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
INFO:causalml:Bootstrap Confidence Intervals for ATE
100%|██████████| 100/100 [01:56<00:00, 1.17s/it]
[53]:
np.vstack((ate_r_lb_b, ate_r_b, ate_r_ub_b))
[53]:
array([[0.37951505],
[0.54612646],
[0.53701368]])
Without Propensity Score Input
[54]:
ate_r_b_no_p, ate_r_lb_b_no_p, ate_r_ub_b_no_p = learner_r.estimate_ate(X=X, treatment=treatment, y=y, bootstrap_ci=True,
n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
INFO:causalml:Bootstrap Confidence Intervals for ATE
100%|██████████| 100/100 [02:42<00:00, 1.63s/it]
[55]:
np.vstack((ate_r_lb_b_no_p, ate_r_b_no_p, ate_r_ub_b_no_p))
[55]:
array([[0.37126915],
[0.50635052],
[0.51400059]])
CATE
With Propensity Score Input
[56]:
learner_r = BaseRRegressor(XGBRegressor(), control_name='control')
cate_r = learner_r.fit_predict(X=X, treatment=treatment, y=y, p=e)
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
[57]:
cate_r
[57]:
array([[ 1.57365084],
[-0.63619554],
[-0.05320793],
...,
[ 0.56346375],
[ 0.56288183],
[ 0.87085617]])
Without Propensity Score Input
[58]:
cate_r_no_p = learner_r.fit_predict(X=X, treatment=treatment, y=y)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
[59]:
cate_r_no_p
[59]:
array([[-0.19582933],
[-0.29006499],
[ 0.46513131],
...,
[ 0.89712083],
[ 0.81002617],
[ 0.82598114]])
CATE w/ Confidence Intervals
With Propensity Score Input
[60]:
learner_r = BaseRRegressor(XGBRegressor(), control_name='control')
cate_r, cate_r_lb, cate_r_ub = learner_r.fit_predict(X=X, treatment=treatment, y=y, p=e, return_ci=True,
n_bootstraps=100, bootstrap_size=1000)
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
INFO:causalml:Bootstrap Confidence Intervals
100%|██████████| 100/100 [00:46<00:00, 2.15it/s]
[61]:
cate_r
[61]:
array([[ 0.43967736],
[-0.27467608],
[-0.36704457],
...,
[ 1.70213294],
[ 0.53581667],
[ 0.67119908]])
[62]:
cate_r_lb
[62]:
array([[-2.36270347],
[-2.10110987],
[-3.33190218],
...,
[-2.25005704],
[-2.08611215],
[-1.89283199]])
[63]:
cate_r_ub
[63]:
array([[3.23361461],
[4.39421365],
[3.95620847],
...,
[3.15905744],
[3.23586204],
[2.31788745]])
Without Propensity Score Input
[64]:
learner_r = BaseRRegressor(XGBRegressor(), control_name='control')
cate_r_no_p, cate_r_lb_no_p, cate_r_ub_no_p = learner_r.fit_predict(X=X, treatment=treatment, y=y, return_ci=True,
n_bootstraps=100, bootstrap_size=1000)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
INFO:causalml:Bootstrap Confidence Intervals
100%|██████████| 100/100 [00:31<00:00, 3.14it/s]
[65]:
cate_r_no_p
[65]:
array([[-0.14972556],
[ 0.18446118],
[ 0.23380044],
...,
[ 0.55917108],
[-0.16540062],
[ 0.62050438]])
[66]:
cate_r_lb_no_p
[66]:
array([[-2.37674593],
[-1.66803797],
[-3.47868801],
...,
[-1.95877534],
[-2.32770172],
[-1.68704787]])
[67]:
cate_r_ub_no_p
[67]:
array([[2.9130644 ],
[3.99895564],
[3.61212277],
...,
[3.174209 ],
[3.38644627],
[2.62858756]])
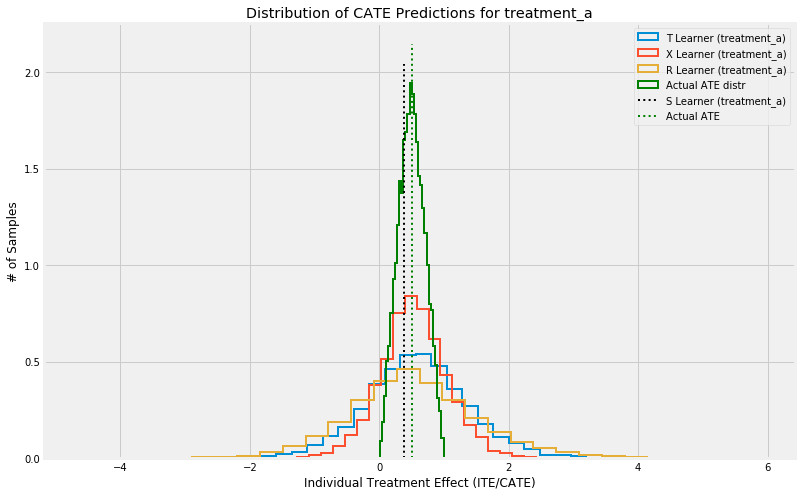
Visualize
[68]:
groups = learner_r._classes
alpha = 1
linewidth = 2
bins = 30
for group,idx in sorted(groups.items(), key=lambda x: x[1]):
plt.figure(figsize=(12,8))
plt.hist(cate_t[:,idx], alpha=alpha, bins=bins, label='T Learner ({})'.format(group),
histtype='step', linewidth=linewidth, density=True)
plt.hist(cate_x[:,idx], alpha=alpha, bins=bins, label='X Learner ({})'.format(group),
histtype='step', linewidth=linewidth, density=True)
plt.hist(cate_r[:,idx], alpha=alpha, bins=bins, label='R Learner ({})'.format(group),
histtype='step', linewidth=linewidth, density=True)
plt.hist(tau, alpha=alpha, bins=bins, label='Actual ATE distr',
histtype='step', linewidth=linewidth, color='green', density=True)
plt.vlines(cate_s[0,idx], 0, plt.axes().get_ylim()[1], label='S Learner ({})'.format(group),
linestyles='dotted', linewidth=linewidth)
plt.vlines(tau.mean(), 0, plt.axes().get_ylim()[1], label='Actual ATE',
linestyles='dotted', linewidth=linewidth, color='green')
plt.title('Distribution of CATE Predictions for {}'.format(group))
plt.xlabel('Individual Treatment Effect (ITE/CATE)')
plt.ylabel('# of Samples')
_=plt.legend()
Multiple Treatment Case
Generate synthetic data
Note: we randomize the assignment of treatment flag AFTER the synthetic data generation process, so it doesn’t make sense to measure accuracy metrics here. Next steps would be to include multi-treatment in the DGP itself.
[69]:
# Generate synthetic data using mode 1
y, X, treatment, tau, b, e = synthetic_data(mode=1, n=10000, p=8, sigma=1.0)
treatment = np.array([('treatment_a' if np.random.random() > 0.2 else 'treatment_b')
if val==1 else 'control' for val in treatment])
e = {group: e for group in np.unique(treatment)}
[70]:
pd.Series(treatment).value_counts()
[70]:
control 4768
treatment_a 4146
treatment_b 1086
dtype: int64
S-Learner
ATE
[71]:
learner_s = BaseSRegressor(XGBRegressor(), control_name='control')
ate_s = learner_s.estimate_ate(X=X, treatment=treatment, y=y, return_ci=False, bootstrap_ci=False)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.6339
INFO:causalml: RMSE (Treatment): 0.6447
INFO:causalml: sMAPE (Control): 0.6148
INFO:causalml: sMAPE (Treatment): 0.3498
INFO:causalml: Gini (Control): 0.8528
INFO:causalml: Gini (Treatment): 0.8492
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.5584
INFO:causalml: RMSE (Treatment): 0.4771
INFO:causalml: sMAPE (Control): 0.5699
INFO:causalml: sMAPE (Treatment): 0.2768
INFO:causalml: Gini (Control): 0.8921
INFO:causalml: Gini (Treatment): 0.9227
[72]:
ate_s
[72]:
array([0.58349553, 0.58778215])
[73]:
learner_s._classes
[73]:
{'treatment_a': 0, 'treatment_b': 1}
ATE w/ Confidence Intervals
[74]:
alpha = 0.05
learner_s = BaseSRegressor(XGBRegressor(), ate_alpha=alpha, control_name='control')
ate_s, ate_s_lb, ate_s_ub = learner_s.estimate_ate(X=X, treatment=treatment, y=y, return_ci=True,
bootstrap_ci=False)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.6339
INFO:causalml: RMSE (Treatment): 0.6447
INFO:causalml: sMAPE (Control): 0.6148
INFO:causalml: sMAPE (Treatment): 0.3498
INFO:causalml: Gini (Control): 0.8528
INFO:causalml: Gini (Treatment): 0.8492
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.5584
INFO:causalml: RMSE (Treatment): 0.4771
INFO:causalml: sMAPE (Control): 0.5699
INFO:causalml: sMAPE (Treatment): 0.2768
INFO:causalml: Gini (Control): 0.8921
INFO:causalml: Gini (Treatment): 0.9227
[75]:
np.vstack((ate_s_lb, ate_s, ate_s_ub))
[75]:
array([[0.5555693 , 0.55278018],
[0.58349553, 0.58778215],
[0.61142176, 0.62278413]])
ATE w/ Boostrap Confidence Intervals
[76]:
ate_s_b, ate_s_lb_b, ate_s_ub_b = learner_s.estimate_ate(X=X, treatment=treatment, y=y, return_ci=True,
bootstrap_ci=True, n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.6339
INFO:causalml: RMSE (Treatment): 0.6447
INFO:causalml: sMAPE (Control): 0.6148
INFO:causalml: sMAPE (Treatment): 0.3498
INFO:causalml: Gini (Control): 0.8528
INFO:causalml: Gini (Treatment): 0.8492
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.5584
INFO:causalml: RMSE (Treatment): 0.4771
INFO:causalml: sMAPE (Control): 0.5699
INFO:causalml: sMAPE (Treatment): 0.2768
INFO:causalml: Gini (Control): 0.8921
INFO:causalml: Gini (Treatment): 0.9227
INFO:causalml:Bootstrap Confidence Intervals for ATE
100%|██████████| 100/100 [01:40<00:00, 1.00s/it]
[77]:
np.vstack((ate_s_lb_b, ate_s_b, ate_s_ub_b))
[77]:
array([[0.52550035, 0.52550035],
[0.58349553, 0.58778215],
[0.64944596, 0.64944596]])
CATE
[78]:
learner_s = BaseSRegressor(XGBRegressor(), control_name='control')
cate_s = learner_s.fit_predict(X=X, treatment=treatment, y=y, return_ci=False)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.6339
INFO:causalml: RMSE (Treatment): 0.6447
INFO:causalml: sMAPE (Control): 0.6148
INFO:causalml: sMAPE (Treatment): 0.3498
INFO:causalml: Gini (Control): 0.8528
INFO:causalml: Gini (Treatment): 0.8492
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.5584
INFO:causalml: RMSE (Treatment): 0.4771
INFO:causalml: sMAPE (Control): 0.5699
INFO:causalml: sMAPE (Treatment): 0.2768
INFO:causalml: Gini (Control): 0.8921
INFO:causalml: Gini (Treatment): 0.9227
[79]:
cate_s
[79]:
array([[ 0.91381967, 0.82956386],
[-0.17692167, -0.15709245],
[ 0.90877771, 0.92332006],
...,
[ 0.86159408, 0.53687155],
[ 0.66541922, 0.78590739],
[ 1.05691028, 1.03345728]])
CATE w/ Confidence Intervals
[80]:
alpha = 0.05
learner_s = BaseSRegressor(XGBRegressor(), ate_alpha=alpha, control_name='control')
cate_s, cate_s_lb, cate_s_ub = learner_s.fit_predict(X=X, treatment=treatment, y=y, return_ci=True,
n_bootstraps=100, bootstrap_size=3000)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.6339
INFO:causalml: RMSE (Treatment): 0.6447
INFO:causalml: sMAPE (Control): 0.6148
INFO:causalml: sMAPE (Treatment): 0.3498
INFO:causalml: Gini (Control): 0.8528
INFO:causalml: Gini (Treatment): 0.8492
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.5584
INFO:causalml: RMSE (Treatment): 0.4771
INFO:causalml: sMAPE (Control): 0.5699
INFO:causalml: sMAPE (Treatment): 0.2768
INFO:causalml: Gini (Control): 0.8921
INFO:causalml: Gini (Treatment): 0.9227
INFO:causalml:Bootstrap Confidence Intervals
100%|██████████| 100/100 [01:03<00:00, 1.58it/s]
[81]:
cate_s
[81]:
array([[ 0.91381967, 0.82956386],
[-0.17692167, -0.15709245],
[ 0.90877771, 0.92332006],
...,
[ 0.86159408, 0.53687155],
[ 0.66541922, 0.78590739],
[ 1.05691028, 1.03345728]])
[82]:
cate_s_lb
[82]:
array([[ 0.23816384, -0.32713253],
[-0.44141183, -0.42676411],
[-0.00206863, -0.43860602],
...,
[ 0.29240462, -0.16563866],
[-0.01797467, -0.10772878],
[-0.51486325, -0.31691882]])
[83]:
cate_s_ub
[83]:
array([[1.40557503, 1.1807412 ],
[1.06860972, 1.55298753],
[1.38529261, 1.6596471 ],
...,
[1.56729684, 1.47052228],
[1.16166003, 1.1144281 ],
[1.68127107, 1.58984778]])
T-Learner
ATE w/ Confidence Intervals
[84]:
learner_t = BaseTRegressor(XGBRegressor(), control_name='control')
ate_t, ate_t_lb, ate_t_ub = learner_t.estimate_ate(X=X, treatment=treatment, y=y)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.4669
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.2675
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9297
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.0747
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.0568
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9984
[85]:
np.vstack((ate_t_lb, ate_t, ate_t_ub))
[85]:
array([[0.53107041, 0.5296616 ],
[0.55739303, 0.55794811],
[0.58371565, 0.58623463]])
ATE w/ Boostrap Confidence Intervals
[86]:
ate_t_b, ate_t_lb_b, ate_t_ub_b = learner_t.estimate_ate(X=X, treatment=treatment, y=y, bootstrap_ci=True,
n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.4669
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.2675
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9297
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.0747
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.0568
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9984
INFO:causalml:Bootstrap Confidence Intervals for ATE
100%|██████████| 100/100 [01:32<00:00, 1.08it/s]
[87]:
np.vstack((ate_t_lb_b, ate_t_b, ate_t_ub_b))
[87]:
array([[0.51777538, 0.51777538],
[0.55739303, 0.55794811],
[0.67471492, 0.67471492]])
CATE
[88]:
learner_t = BaseTRegressor(XGBRegressor(), control_name='control')
cate_t = learner_t.fit_predict(X=X, treatment=treatment, y=y)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.4669
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.2675
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9297
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.0747
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.0568
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9984
[89]:
cate_t
[89]:
array([[ 1.47525787, -0.06651461],
[ 1.26169336, 1.14718354],
[ 1.68760026, 0.75878632],
...,
[ 0.37292147, 0.20537615],
[ 0.84290075, 0.80045319],
[ 1.64227223, 1.91352534]])
CATE w/ Confidence Intervals
[90]:
learner_t = BaseTRegressor(XGBRegressor(), control_name='control')
cate_t, cate_t_lb, cate_t_ub = learner_t.fit_predict(X=X, treatment=treatment, y=y, return_ci=True, n_bootstraps=100,
bootstrap_size=3000)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.4669
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.2675
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9297
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.0747
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.0568
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9984
INFO:causalml:Bootstrap Confidence Intervals
100%|██████████| 100/100 [01:06<00:00, 1.51it/s]
[91]:
cate_t
[91]:
array([[ 1.47525787, -0.06651461],
[ 1.26169336, 1.14718354],
[ 1.68760026, 0.75878632],
...,
[ 0.37292147, 0.20537615],
[ 0.84290075, 0.80045319],
[ 1.64227223, 1.91352534]])
[92]:
cate_t_lb
[92]:
array([[-0.18706408, -0.84940575],
[-1.01419897, -0.7311732 ],
[-0.0427315 , -0.16378173],
...,
[-0.39076423, -0.16869925],
[-0.17401927, -0.19503389],
[-0.61903974, -1.15808628]])
[93]:
cate_t_ub
[93]:
array([[2.47563672, 1.69891493],
[2.04089584, 1.76605188],
[2.3567108 , 2.40833322],
...,
[2.17926003, 2.26919731],
[2.15714553, 1.91076722],
[2.27031788, 2.03901908]])
X-Learner
ATE w/ Confidence Intervals
With Propensity Score Input
[94]:
learner_x = BaseXRegressor(XGBRegressor(), control_name='control')
ate_x, ate_x_lb, ate_x_ub = learner_x.estimate_ate(X=X, treatment=treatment, y=y, p=e)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.4669
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.2675
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9297
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.0747
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.0568
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9984
[95]:
np.vstack((ate_x_lb, ate_x, ate_x_ub))
[95]:
array([[0.49573269, 0.54002602],
[0.51860246, 0.56163457],
[0.54147223, 0.58324311]])
Without Propensity Score Input
[96]:
ate_x_no_p, ate_x_lb_no_p, ate_x_ub_no_p = learner_x.estimate_ate(X=X, treatment=treatment, y=y)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.4669
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.2675
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9297
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.0747
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.0568
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9984
[97]:
np.vstack((ate_x_lb_no_p, ate_x_no_p, ate_x_ub_no_p))
[97]:
array([[0.50418298, 0.56976992],
[0.52706595, 0.59243233],
[0.54994892, 0.61509475]])
ATE w/ Boostrap Confidence Intervals
With Propensity Score Input
[98]:
ate_x_b, ate_x_lb_b, ate_x_ub_b = learner_x.estimate_ate(X=X, treatment=treatment, y=y, p=e, bootstrap_ci=True,
n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.4669
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.2675
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9297
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.0747
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.0568
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9984
INFO:causalml:Bootstrap Confidence Intervals for ATE
100%|██████████| 100/100 [02:55<00:00, 1.75s/it]
[99]:
np.vstack((ate_x_lb_b, ate_x_b, ate_x_ub_b))
[99]:
array([[0.49600789, 0.49600789],
[0.51860246, 0.56163457],
[0.63696386, 0.63696386]])
Without Propensity Score Input
[100]:
ate_x_b_no_p, ate_x_lb_b_no_p, ate_x_ub_b_no_p = learner_x.estimate_ate(X=X, treatment=treatment, y=y, bootstrap_ci=True,
n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.4669
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.2675
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9297
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.0747
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.0568
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9984
INFO:causalml:Bootstrap Confidence Intervals for ATE
100%|██████████| 100/100 [02:54<00:00, 1.74s/it]
[101]:
np.vstack((ate_x_lb_b_no_p, ate_x_b_no_p, ate_x_ub_b_no_p))
[101]:
array([[0.50100288, 0.50100288],
[0.52706414, 0.59242806],
[0.66020792, 0.66020792]])
CATE
With Propensity Score Input
[102]:
learner_x = BaseXRegressor(XGBRegressor(), control_name='control')
cate_x = learner_x.fit_predict(X=X, treatment=treatment, y=y, p=e)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.4669
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.2675
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9297
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.0747
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.0568
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9984
[103]:
cate_x
[103]:
array([[ 0.57149441, 0.10240081],
[-0.43192272, 1.48913118],
[ 1.13622262, 0.65923928],
...,
[ 0.44651704, -0.23119723],
[ 0.93875551, 0.77003003],
[ 0.96697381, 0.99990004]])
Without Propensity Score Input
[104]:
cate_x_no_p = learner_x.fit_predict(X=X, treatment=treatment, y=y)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.4669
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.2675
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9297
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.0747
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.0568
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9984
[105]:
cate_x_no_p
[105]:
array([[ 0.62959351, -0.00493521],
[-0.48863166, 1.54109948],
[ 1.17988308, 1.26200671],
...,
[ 0.41320951, 0.73251634],
[ 0.91104634, 0.82359481],
[ 1.08867931, 1.44193089]])
CATE w/ Confidence Intervals
With Propensity Score Input
[106]:
learner_x = BaseXRegressor(XGBRegressor(), control_name='control')
cate_x, cate_x_lb, cate_x_ub = learner_x.fit_predict(X=X, treatment=treatment, y=y, p=e, return_ci=True,
n_bootstraps=100, bootstrap_size=1000)
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.4669
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.2675
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9297
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.0747
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.0568
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9984
INFO:causalml:Bootstrap Confidence Intervals
100%|██████████| 100/100 [00:51<00:00, 1.94it/s]
[107]:
learner_x._classes
[107]:
{'treatment_a': 0, 'treatment_b': 1}
[108]:
cate_x
[108]:
array([[ 0.57149441, 0.10240081],
[-0.43192272, 1.48913118],
[ 1.13622262, 0.65923928],
...,
[ 0.44651704, -0.23119723],
[ 0.93875551, 0.77003003],
[ 0.96697381, 0.99990004]])
[109]:
cate_x_lb
[109]:
array([[-0.23574115, -0.21029023],
[-0.95699419, -1.05203708],
[-0.49402807, -0.48280283],
...,
[-0.12162789, -0.26408791],
[-0.52562958, -0.19338615],
[-0.40858565, -0.88119588]])
[110]:
cate_x_ub
[110]:
array([[1.79950407, 2.11258332],
[1.45309225, 1.48831446],
[1.75564219, 2.03222137],
...,
[2.15191078, 2.30032378],
[1.65228261, 1.40411322],
[1.74815254, 1.68257617]])
Without Propensity Score Input
[111]:
cate_x_no_p, cate_x_lb_no_p, cate_x_ub_no_p = learner_x.fit_predict(X=X, treatment=treatment, y=y, return_ci=True,
n_bootstraps=100, bootstrap_size=1000)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Error metrics for group treatment_a
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.4669
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.2675
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9297
INFO:causalml:Error metrics for group treatment_b
INFO:causalml: RMSE (Control): 0.4743
INFO:causalml: RMSE (Treatment): 0.0747
INFO:causalml: sMAPE (Control): 0.5062
INFO:causalml: sMAPE (Treatment): 0.0568
INFO:causalml: Gini (Control): 0.9280
INFO:causalml: Gini (Treatment): 0.9984
INFO:causalml:Bootstrap Confidence Intervals
100%|██████████| 100/100 [00:51<00:00, 1.94it/s]
[112]:
learner_x._classes
[112]:
{'treatment_a': 0, 'treatment_b': 1}
[113]:
cate_x_no_p
[113]:
array([[ 0.6294132 , -0.00492528],
[-0.48876998, 1.54111376],
[ 1.17989094, 1.2620318 ],
...,
[ 0.41319463, 0.73237091],
[ 0.9108665 , 0.82359564],
[ 1.08868219, 1.441931 ]])
[114]:
cate_x_lb_no_p
[114]:
array([[-0.10073893, -0.38800051],
[-0.81971717, -0.8298923 ],
[-0.18606629, -0.32586878],
...,
[ 0.18372251, -0.12170252],
[-0.21309623, -0.38600234],
[-0.44863794, -0.39716903]])
[115]:
cate_x_ub_no_p
[115]:
array([[2.00312255, 2.10486085],
[1.59355675, 1.76340695],
[1.77980204, 2.35535097],
...,
[1.94828429, 1.94720835],
[2.04021647, 1.71337955],
[1.60121219, 1.82820234]])
R-Learner
ATE w/ Confidence Intervals
With Propensity Score Input
[116]:
learner_r = BaseRRegressor(XGBRegressor(), control_name='control')
ate_r, ate_r_lb, ate_r_ub = learner_r.estimate_ate(X=X, treatment=treatment, y=y, p=e)
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
INFO:causalml:training the treatment effect model for treatment_b with R-loss
[117]:
np.vstack((ate_r_lb, ate_r, ate_r_ub))
[117]:
array([[0.52326968, 0.57744164],
[0.52374892, 0.5781462 ],
[0.52422816, 0.57885076]])
Without Propensity Score Input
[118]:
learner_r = BaseRRegressor(XGBRegressor(), control_name='control')
ate_r_no_p, ate_r_lb_no_p, ate_r_ub_no_p = learner_r.estimate_ate(X=X, treatment=treatment, y=y)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Calibrating propensity scores.
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
INFO:causalml:training the treatment effect model for treatment_b with R-loss
[119]:
np.vstack((ate_r_lb_no_p, ate_r_no_p, ate_r_ub_no_p))
[119]:
array([[0.44161159, 0.71836119],
[0.44209269, 0.71904979],
[0.44257378, 0.71973838]])
[120]:
learner_r.propensity_model
[120]:
{'treatment_a': {'all training': LogisticRegressionCV(Cs=array([1.00230524, 2.15608891, 4.63802765, 9.97700064]),
class_weight=None,
cv=KFold(n_splits=5, random_state=None, shuffle=True),
dual=False, fit_intercept=True, intercept_scaling=1.0,
l1_ratios=array([0.001 , 0.33366667, 0.66633333, 0.999 ]),
max_iter=100, multi_class='auto', n_jobs=None,
penalty='elasticnet', random_state=None, refit=True,
scoring=None, solver='saga', tol=0.0001, verbose=0)},
'treatment_b': {'all training': LogisticRegressionCV(Cs=array([1.00230524, 2.15608891, 4.63802765, 9.97700064]),
class_weight=None,
cv=KFold(n_splits=5, random_state=None, shuffle=True),
dual=False, fit_intercept=True, intercept_scaling=1.0,
l1_ratios=array([0.001 , 0.33366667, 0.66633333, 0.999 ]),
max_iter=100, multi_class='auto', n_jobs=None,
penalty='elasticnet', random_state=None, refit=True,
scoring=None, solver='saga', tol=0.0001, verbose=0)}}
ATE w/ Boostrap Confidence Intervals
With Propensity Score Input
[121]:
ate_r_b, ate_r_lb_b, ate_r_ub_b = learner_r.estimate_ate(X=X, treatment=treatment, y=y, p=e, bootstrap_ci=True,
n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
INFO:causalml:training the treatment effect model for treatment_b with R-loss
INFO:causalml:Bootstrap Confidence Intervals for ATE
100%|██████████| 100/100 [02:19<00:00, 1.39s/it]
[122]:
np.vstack((ate_r_lb_b, ate_r_b, ate_r_ub_b))
[122]:
array([[0.40326436, 0.40326436],
[0.50620059, 0.5478152 ],
[0.5697328 , 0.5697328 ]])
Without Propensity Score Input
[123]:
learner_r = BaseRRegressor(XGBRegressor(), control_name='control')
ate_r_b_no_p, ate_r_lb_b_no_p, ate_r_ub_b_no_p = learner_r.estimate_ate(X=X, treatment=treatment, y=y, bootstrap_ci=True,
n_bootstraps=100, bootstrap_size=5000)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Calibrating propensity scores.
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
INFO:causalml:training the treatment effect model for treatment_b with R-loss
INFO:causalml:Bootstrap Confidence Intervals for ATE
100%|██████████| 100/100 [02:19<00:00, 1.39s/it]
[124]:
np.vstack((ate_r_lb_b_no_p, ate_r_b_no_p, ate_r_ub_b_no_p))
[124]:
array([[0.45994051, 0.45994051],
[0.44481491, 0.66323246],
[0.68981572, 0.68981572]])
CATE
With Propensity Score Input
[125]:
learner_r = BaseRRegressor(XGBRegressor(), control_name='control')
cate_r = learner_r.fit_predict(X=X, treatment=treatment, y=y, p=e)
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
INFO:causalml:training the treatment effect model for treatment_b with R-loss
[126]:
cate_r
[126]:
array([[ 5.57098567e-01, 1.77359581e-03],
[ 1.08587885e+00, 2.48472750e-01],
[ 3.34437251e-01, 1.69020355e+00],
...,
[-9.96065974e-01, -8.98482800e-02],
[ 1.70625651e+00, 9.55640435e-01],
[-1.88456130e+00, 6.50659442e-01]])
Without Propensity Score Input
[127]:
learner_r = BaseRRegressor(XGBRegressor(), control_name='control')
cate_r_no_p = learner_r.fit_predict(X=X, treatment=treatment, y=y)
INFO:causalml:Generating propensity score
INFO:causalml:Calibrating propensity scores.
INFO:causalml:Calibrating propensity scores.
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
INFO:causalml:training the treatment effect model for treatment_b with R-loss
[128]:
cate_r_no_p
[128]:
array([[ 0.55478877, 0.87992519],
[ 1.10120189, 1.29564619],
[ 0.62448621, 0.41555083],
...,
[-0.53886592, 0.44593787],
[ 1.25231111, 0.79904991],
[-0.64419305, -0.23014426]])
CATE w/ Confidence Intervals
With Propensity Score Input
[129]:
learner_r = BaseRRegressor(XGBRegressor(), control_name='control')
cate_r, cate_r_lb, cate_r_ub = learner_r.fit_predict(X=X, treatment=treatment, y=y, p=e, return_ci=True,
n_bootstraps=100, bootstrap_size=1000)
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
INFO:causalml:training the treatment effect model for treatment_b with R-loss
INFO:causalml:Bootstrap Confidence Intervals
100%|██████████| 100/100 [00:37<00:00, 2.65it/s]
[130]:
cate_r
[130]:
array([[ 1.75007784, 0.67752302],
[ 0.77257723, 0.12910607],
[ 1.08854032, 0.81679094],
...,
[-0.92310214, 0.645491 ],
[ 0.92478108, 0.79903334],
[-0.48311949, 1.00291944]])
[131]:
cate_r_lb
[131]:
array([[-0.801657 , -0.48754777],
[-3.05317249, -5.37572038],
[-1.50823961, -1.16822439],
...,
[-1.27909884, -1.2460175 ],
[-1.42656819, -1.59059022],
[-1.90115855, -2.10247456]])
[132]:
cate_r_ub
[132]:
array([[4.06750882, 3.68516954],
[4.21587243, 4.50271177],
[4.33370841, 3.79358828],
...,
[3.53610538, 3.48638564],
[3.71832166, 3.48292163],
[5.01262635, 3.27047309]])
Without Propensity Score Input
[ ]:
learner_r = BaseRRegressor(XGBRegressor(), control_name='control')
cate_r_no_p, cate_r_lb_no_p, cate_r_ub_no_p = learner_r.fit_predict(X=X, treatment=treatment, y=y, p=e, return_ci=True,
n_bootstraps=100, bootstrap_size=1000)
INFO:causalml:generating out-of-fold CV outcome estimates
INFO:causalml:training the treatment effect model for treatment_a with R-loss
INFO:causalml:training the treatment effect model for treatment_b with R-loss
INFO:causalml:Bootstrap Confidence Intervals
2%|▏ | 2/100 [00:00<00:36, 2.69it/s]
[ ]:
cate_r_no_p
[ ]:
cate_r_lb_no_p
[ ]:
cate_r_ub_no_p